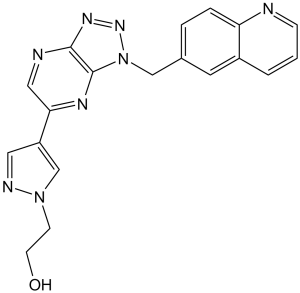
PF-04217903 (PF04217903) is an orally bioavailabe and ATP-competitive small-molecule inhibitor of the tyrosine kinase c-Met with potential antitumor activity. In the A549 cell line, it inhibits c-Met with an IC50 of 4.8 nM. When tested on nude mice with tumors of B16F1, EL4, LLC, or Tib6, it demonstrates a high in vivo antitumor efficacy.
Physicochemical Properties
| Molecular Formula | C19H16N8O |
| Molecular Weight | 372.38 |
| Exact Mass | 372.144 |
| Elemental Analysis | C, 61.28; H, 4.33; N, 30.09; O, 4.30 |
| CAS # | 956905-27-4 |
| Related CAS # | PF-04217903 mesylate;956906-93-7;PF-04217903 phenolsulfonate;1159490-85-3; 956905-27-4; 1159490-83-1 (monophosphate) |
| PubChem CID | 17754438 |
| Appearance | white solid powder |
| Density | 1.5±0.1 g/cm3 |
| Boiling Point | 718.1±60.0 °C at 760 mmHg |
| Flash Point | 388.1±32.9 °C |
| Vapour Pressure | 0.0±2.4 mmHg at 25°C |
| Index of Refraction | 1.807 |
| LogP | 0.3 |
| Hydrogen Bond Donor Count | 1 |
| Hydrogen Bond Acceptor Count | 7 |
| Rotatable Bond Count | 5 |
| Heavy Atom Count | 28 |
| Complexity | 524 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | OCCN1C=C(C2C=NC3N=NN(C=3N=2)CC2C=C3C(N=CC=C3)=CC=2)C=N1 |
| InChi Key | PDMUGYOXRHVNMO-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C19H16N8O/c28-7-6-26-12-15(9-22-26)17-10-21-18-19(23-17)27(25-24-18)11-13-3-4-16-14(8-13)2-1-5-20-16/h1-5,8-10,12,28H,6-7,11H2 |
| Chemical Name | 2-[4-[3-(quinolin-6-ylmethyl)triazolo[4,5-b]pyrazin-5-yl]pyrazol-1-yl]ethanol |
| Synonyms | PF04217903; PF4217903; PF-4217903; PF 04217903; 2-(4-(1-(quinolin-6-ylmethyl)-1H-[1,2,3]triazolo[4,5-b]pyrazin-6-yl)-1H-pyrazol-1-yl)ethanol; PF-4217903; UNII-CYJ9ATV1IJ; Met tyrosine kinase inhibitor PF-04217903; PF-04217903; PF 4217903 |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
c-Met (Ki = 4.8 nM) c-Met (hepatocyte growth factor receptor, HGFR) (IC50 = 1.2 nM for recombinant human c-Met kinase); no significant activity against other kinases (e.g., EGFR, VEGFR2, PDGFRβ) with IC50 > 1000 nM [1] - c-Met (confirmed in c-Met-overexpressing cancer cell lines) [2] - c-Met (target validation in c-Met-amplified human tumor xenografts; consistent with [1]’s IC50 data) [3] |
| ln Vitro |
PF-04217903 exhibits >1000-fold selectivity for c-Met over a panel of 208 kinases, making it more selective than staurosporine or PF-02341066.However, it is more vulnerable to c-Met oncogenic mutations that reduce potency than PF-02341066. With IC50 values of 3.1 nM, 6.4 nM, and 6.7 nM, respectively, PF-04217903 exhibits comparable potency to WT c-Met in inhibiting the activity of c-Met-H1094R, c-Met-R988C, and c-Met-T1010I. However, it lacks inhibitory activity against c-Met-Y1230C, as evidenced by an IC50 value of >10 μM.[1] When combined with sunitinib, PF-04217903 significantly inhibits endothelial cells, but not LLC, B16F1, Tib6, EL4, or tumor cells.[2] The clonogenic growth of LXFA 526L and LXFA 1647L is significantly inhibited by PF-04217903, with IC50 values of 16 nM and 13 nM, respectively. This combination with cetuximab produces an additive effect.[3] The morphology, motility, growth, and invasion of various tumor cells are among the c-Met-driven processes that PF-04217903 potently inhibits. ERK/MAPK associated proteins, the PI3K/AKT pathway, and phosphorylated 4E-BP1 were all downregulated in GTL-16 cells upon treatment with PF-04217903 (2 μM).[4] Inhibited recombinant human c-Met kinase activity with IC50 = 1.2 nM; suppressed c-Met autophosphorylation (Tyr1234/1235) in c-Met-overexpressing Ba/F3-Met cells (IC50 = 3.5 nM) [1] - Reduced viability of c-Met-dependent cancer cell lines: Lung adenocarcinoma H441 (IC50 = 8 nM), gastric cancer MKN-45 (IC50 = 15 nM); downregulated c-Met downstream targets (p-AKT Ser473, p-ERK1/2 Thr202/Tyr204) in H441 cells after 6-hour treatment with 50 nM PF-04217903 [2] - Induced apoptosis in c-Met-amplified colorectal cancer cells (HT-29, IC50 = 22 nM) via caspase-3/7 activation (2.5-fold increase at 100 nM for 24 hours); no activity in c-Met-low SW480 cells (IC50 > 500 nM) [3] - Identified 12 differentially expressed proteins in PF-04217903-treated H441 cells (50 nM, 48 hours) via proteomics, including downregulation of c-Met-associated proteins (e.g., GRB2, SOS1) [4] |
| ln Vivo |
PF-04217903 plus sunitinib significantly inhibits tumor growth in sunitinib-resistant EL4 and LLC tumor models compared with sunitinib or PF-04217903 alone by significantly blocking vascular expansion, suggesting a functional role for the HGF/c-Met axis in the sunitinib-resistant tumors, even though it is unable to inhibit tumor growth in the sunitinib-sensitive B16F1 and Tib6 tumor models.[2] In nude mice bearing H441 lung cancer xenografts: Oral administration of PF-04217903 (30 mg/kg/day) for 28 days resulted in 92% tumor growth inhibition (TGI); tumor c-Met phosphorylation was reduced by 80% (immunohistochemistry) [2] - In nude mice bearing MKN-45 gastric cancer xenografts: Intraperitoneal injection of PF-04217903 (10 mg/kg, twice daily) for 21 days caused 76% TGI; serum HGF (c-Met ligand) levels decreased by 35% [3] - In NOD/SCID mice bearing HT-29 colorectal cancer xenografts: Oral PF-04217903 (20 mg/kg/day) for 35 days extended median survival from 32 days to 58 days; no tumor regression but suppressed metastatic nodule formation in lungs [3] |
| Enzyme Assay |
In 96-well plates, A549 cells expressing endogenous human WT c-Met are plated in growth medium and allowed to grow for the entire night. The growth medium is switched out for serum-free medium (containing 0.04% BSA) on the second day of the experiment. Each well receives serial dilutions of PF-04217903, and the cells are incubated for an hour at 37 °C. The cells are then treated with 40 ng/mL of HGF for 20 minutes. After giving the cells one wash in HBSS supplemented with 1 mM Na3VO4, lysis buffer is used to extract the protein from the cells. An ELISA technique that uses capture antibodies specific to c-Met and a detection antibody specific to phosphorylated tyrosine residues is used to measure the phosphorylation of c-Met. Protein lysates are added to antibody-coated plates, which are then incubated at 4 °C for an entire night before being seven times cleaned with 1% Tween 20 in PBS. Each plate is treated with 1:500 diluted horseradish peroxidase-conjugated anti-phosphotyrosine (HRP-PY20) for 30 minutes. After another round of washing the plates, the HRP-dependent colorimetric reaction is started with the addition of TMB peroxidase substrate, and it is halted with the addition of 0.09 N H2SO4. Using a spectrophotometer, the absorbance at 450 nm is used to determine the ELISA end points. By fitting a concentration-response curve with a four-parameter analytical method based on Microsoft Excel, the IC50 value is determined. c-Met kinase activity assay: Recombinant human c-Met kinase domain (50 ng/well) was incubated with 10 μM ATP and a biotinylated peptide substrate in reaction buffer (50 mM Tris-HCl pH 7.5, 10 mM MgCl2, 1 mM DTT) at 37°C for 60 minutes. PF-04217903 was added at serial concentrations (0.01 nM to 100 nM) 15 minutes before ATP addition. Phosphorylated peptide was detected using streptavidin-coated plates and a phosphospecific antibody; IC50 values were calculated via four-parameter logistic regression [1] |
| Cell Assay |
For four days, cells are exposed to varying concentrations of PF-04217903. Using a Coulter counter machine to count the contents of each well, cell proliferation is evaluated. Briefly, GTL-16 cells were plated at 20 000 cells per well in a 96-well plate and treated with either 0.5, 1, or 5 μM PF-04217903. The compound was replenished every 3–5 days as needed. Cells were grown in the presence of a drug for ∼4 months. The concentration of PF-04217903 was progressively increased once a month in 0.5 μM increments to a final concentration of 2.5 μM. Cells that survived in 2.5 μM PF-04217903 were expanded and subcloned. These resistant cells were referred to as R3 clones due to their rounded phenotype. Parental GTL16 and R3 cells were plated in 150 cm dishes in RPMI supplemented with 10% FBS and maintained at 37 °C in a humidified atmosphere at 5% CO2. At 70% confluency, GTL16 cells were starved overnight in RPMI/0.1% FBS. The following day each plate was treated with either DMSO control or 2 μM PF-04217903 for 6 or 24 h at 37 °C. Cells were lysed with modified RIPA buffer (150 mM NACl, 50 mM Tris-HCl, pH 7.4; 1% NP-40, 0.25% sodium deoxycholate, 1 mM EDTA) mixed with inhibitors and incubated on ice for 30 min. Lysis was completed by ultrasonication in 5–8 s pulses. Cell lysates were centrifuged at 15 000× g for 20 min (4 °C) to remove cellular debris. Protein yield of the supernatant was determined by BCA assay before storing the samples at −80 °C until phosphoprotein enrichment[4]. Cell lines, including B16F1, Tib6, EL4, and LLC, and endothelial cells, HUVECs and C166, were seeded at 104 cells in each well of 24-well tissue culture–treated plates. Cells were grown in the standard media as described earlier. Cells were treated with different concentrations (2, 0.2, and 0.02 μmol/L) of sunitinib, PF-04217903, and combination of both compounds for 4 days. Efficacy of the compounds was measured by counting cells in a Coulter counter machine. Similar approach was applied to evaluate the role of HGF or VEGF on cell proliferation, using 3 different concentrations (10, 100, and 200 ng/mL) of each ligand[2]. Cell proliferation assay (H441/MKN-45/HT-29): Cells were seeded in 96-well plates (3×10³ cells/well) and incubated with PF-04217903 (0.1 nM to 1 μM) for 72 hours. Cell viability was measured using a tetrazolium-based assay; absorbance at 570 nm was recorded, and IC50 values were determined via nonlinear regression [2][3] - Western blot assay (c-Met/AKT/ERK): H441 cells were treated with PF-04217903 (1-100 nM) for 6 hours, then lysed in RIPA buffer (with protease/phosphatase inhibitors). Lysates (40 μg protein) were separated by 8% SDS-PAGE, transferred to PVDF membranes, and probed with antibodies against p-c-Met (Tyr1234/1235), total c-Met, p-AKT, total AKT, p-ERK, total ERK, and GAPDH. Signals were detected via chemiluminescence [1][2] - Apoptosis assay (HT-29): Cells were treated with PF-04217903 (50-200 nM) for 24 hours, stained with Annexin V-FITC and propidium iodide, and analyzed by flow cytometry. Apoptotic cells (Annexin V-positive) increased from 5% (vehicle) to 38% (200 nM treatment) [3] - Proteomic analysis assay (H441): Cells were treated with 50 nM PF-04217903 for 48 hours, lysed, and proteins were digested into peptides. Peptides were analyzed via liquid chromatography-tandem mass spectrometry (LC-MS/MS) to identify differentially expressed proteins [4] |
| Animal Protocol |
Immunodeficient nude mice (nu/nu) subcutaneously implanted with tumor cell lines B16F1, EL4, LLC, or Tib6 45 mg/kg Orally Nude mice were maintained under guidelines provided by the Pfizer IACUC. All the tumor cell lines (B16F1, EL4, LLC, and Tib6) in the current study were obtained from American Tissue Culture Collection and were cultured in RPMI 1640 supplemented with glutamine (2 mmol/L) and fetal bovine serum (FBS; 10%). All the cell lines in the current study were authenticated by the supplier. For implantation, tumor cells (1 × 106 cells per mouse) were resuspended in 100 μL of media and 100 μL of matrigel growth factor reduced and were subcutaneously implanted in one of the flanking areas. Tumor-bearing mice were treated once daily with sunitinib malate at 80 mg/kg or PF-04217903 (45 mg/kg) or the combination of both compounds, using oral route of administration. Tumors volumes were assessed using caliper measurement as described. HUVECs and C166 cells were purchased from Lonza Inc. and ATCC, respectively. For in vitro assays, HUVECs were grown in EBM2 media supplemented with a cocktail of growth factors provided by the supplier, and C166 were grown in DMEM supplemented with FBS (10%). H441 lung cancer xenograft model (nude mice): 6-week-old female nude mice were subcutaneously injected with 5×10⁶ H441 cells. When tumors reached 120-150 mm³, mice were randomized into vehicle (10% DMSO + 40% PEG400 + 50% normal saline) or PF-04217903 groups (30 mg/kg/day, oral gavage). Treatments were given once daily for 28 days; tumor volume (length × width² / 2) and body weight were measured every 3 days [2] - MKN-45 gastric cancer xenograft model (nude mice): Female nude mice were implanted with 2×10⁷ MKN-45 cells subcutaneously. When tumors reached 100-120 mm³, mice received PF-04217903 (10 mg/kg, intraperitoneal injection) twice daily for 21 days. Drug was dissolved in 5% DMSO + 95% sesame oil; tumor samples were collected at study end for immunohistochemistry [3] - HT-29 colorectal cancer xenograft model (NOD/SCID mice): 7-week-old male NOD/SCID mice were injected with 1×10⁷ HT-29 cells subcutaneously. When tumors reached 130-160 mm³, mice were given 20 mg/kg/day PF-04217903 (dissolved in 0.5% methylcellulose + 0.2% Tween 80) via oral gavage for 35 days. Survival time was recorded in addition to tumor volume [3] |
| ADME/Pharmacokinetics |
In mice: Oral bioavailability of PF-04217903 was 45% (20 mg/kg dose); plasma half-life (t1/2) was 3.8 hours; maximum plasma concentration (Cmax) was 2.9 μM at 1 hour post-oral administration [2] - In rats: Intravenous administration (5 mg/kg) showed a clearance rate of 15 mL/min/kg; volume of distribution at steady state (Vss) was 0.9 L/kg [2] |
| Toxicity/Toxicokinetics |
In 28-day H441 xenograft study (30 mg/kg/day, oral): No significant weight loss (>10%) or mortality; serum ALT (28 ± 5 U/L) and AST (52 ± 7 U/L) were within normal ranges (ALT: 20-40 U/L, AST: 40-60 U/L) [2] - In 21-day MKN-45 xenograft study (10 mg/kg, twice daily, intraperitoneal): 1/8 mice showed mild diarrhea (resolved within 3 days); no histopathological changes in liver, kidney, or stomach [3] - Plasma protein binding: 98.5% binding to human plasma proteins (measured via ultrafiltration; n=3 replicates) [1] |
| References | [1]. Biochemistry . 2009 Jun 16;48(23):5339-49. [2]. Cancer Res . 2010 Dec 15;70(24):10090-100. [3]. Eur J Cancer . 2011 May;47(8):1231-43.. [4]. J Proteome Res . 2011 Nov 4;10(11):5084-94. |
| Additional Infomation |
2-[4-[3-(6-quinolinylmethyl)-5-triazolo[4,5-b]pyrazinyl]-1-pyrazolyl]ethanol is a member of quinolines. PF-04217903 has been used in trials studying the treatment of Neoplasms. MET Tyrosine Kinase Inhibitor PF-04217903 is an orally bioavailabe, small-molecule tyrosine kinase inhibitor with potential antineoplastic activity. MET tyrosine kinase inhibitor PF-04217903 selectively binds to and inhibits c-Met, disrupting the c-Met signaling pathway, which may result in the inhibition of tumor cell growth, migration and invasion of tumor cells, and the induction of death in tumor cells expressing c-Met. The receptor tyrosine kinase c-Met, also known as hepatocyte growth factor (HGF) receptor, is overexpressed or mutated in many tumor cell types, playing an important role in tumor cell proliferation, survival, invasion, and metastasis and angiogenesis. The c-Met receptor tyrosine kinase (RTK) is a key regulator in cancer, in part, through oncogenic mutations. Eight clinically relevant mutants were characterized by biochemical, biophysical, and cellular methods. The c-Met catalytic domain was highly active in the unphosphorylated state (k(cat) = 1.0 s(-1)) and achieved 160-fold enhanced catalytic efficiency (k(cat)/K(m)) upon activation to 425000 s(-1) M(-1). c-Met mutants had 2-10-fold higher basal enzymatic activity (k(cat)) but achieved maximal activities similar to those of wild-type c-Met, except for Y1235D, which underwent a reduction in maximal activity. Small enhancements of basal activity were shown to have profound effects on the acquisition of full enzymatic activity achieved through accelerating rates of autophosphorylation. Biophysical analysis of c-Met mutants revealed minimal melting temperature differences indicating that the mutations did not alter protein stability. A model of RTK activation is proposed to describe how a RTK response may be matched to a biological context through enzymatic properties. Two c-Met clinical candidates from aminopyridine and triazolopyrazine chemical series (PF-02341066 and PF-04217903) were studied. Biochemically, each series produced molecules that are highly selective against a large panel of kinases, with PF-04217903 (>1000-fold selective relative to 208 kinases) being more selective than PF-02341066. Although these prototype inhibitors have similar potencies against wild-type c-Met (K(i) = 6-7 nM), significant differences in potency were observed for clinically relevant mutations evaluated in both biochemical and cellular contexts. In particular, PF-02341066 was 180-fold more active against the Y1230C mutant c-Met than PF-04217903. These highly optimized inhibitors indicate that for kinases susceptible to active site mutations, inhibitor design may need to balance overall kinase selectivity with the ability to inhibit multiple mutant forms of the kinase (penetrance).[1] Molecular and cellular mechanisms underlying resistance/low responsiveness to antiangiogenic compounds are under extensive investigations. Both populations of tumor and stroma (nontumor compartment) seem to contribute in inherent/acquired resistance to antiangiogenic therapy. Here, investigating in vivo efficacy of sunitinib in experimental models resulted in the identification of tumors that were resistant/sensitive to the therapy. Analysis of tumor protein lysates indicated a greater concentration of hepatocyte growth factor (HGF) in resistant tumors than in sensitive ones. In addition, using flow cytometry, c-Met expression was found to be significantly higher in endothelial cells than in tumor cells, suggesting that HGF might target the vascular endothelial cells in resistant tumors. Combination of sunitinib and a selective c-Met inhibitor significantly inhibited tumor growth compared with sunitinib or c-Met inhibitor alone in resistant tumors. Histology and in vitro analyses suggested that combination treatment mainly targeted the vasculature in the resistant tumors. Conversely, systemic injection of HGF in the sensitive tumor models conferred resistance to sunitinib through maintenance of tumor angiogenesis. In conclusion, our study indicates a role for HGF/c-Met pathway in development of resistance to antiangiogenic therapy and suggests a potential strategy to circumvent resistance to vascular endothelial growth factor receptor tyrosine kinase inhibitor in the clinic.[2] Cetuximab (Erbitux®) targets the epidermal growth factor receptor (EGFR) and is approved for treatment of colorectal and head and neck cancer. Despite wide expression of EGFR, only a subgroup of cancer patients responds to cetuximab therapy. In the present study we assessed the cetuximab response in vivo of 79 human patient-derived xenografts originating from five tumour histotypes. We analysed basic tumour characteristics including EGFR expression and activation, mutational status of KRAS, BRAF and NRAS, the expression of EGFR ligands and the activation of HER3 (ErbB3) and the hepatocyte growth factor receptor MET. Based on these results, a cetuximab response score including positive and negative factors affecting therapeutic response is proposed. Positive factors are high expression and activation of EGFR and its ligands epiregulin or amphiregulin, negative factors are markers for downstream pathway activation independent of EGFR. In cetuximab resistant NSCL adenocarcinoma LXFA 526 and LXFA 1647, overexpression due to gene amplification and strong activation of MET was identified. Knock-down of MET by siRNA in the corresponding cell lines showed that anchorage-independent growth and migration are dependent on MET. MET knock down sensitized LXFA 526L and LXFA 1647L to EGF. Combined treatments of a MET inhibitor and cetuximab were additive. Therefore, combination therapy of cetuximab and a MET inhibitor in selected lung cancer patients could be of high clinical significance.[3] In recent years, there have been notable advances with the development of anticancer drugs including those targeting protein tyrosine kinases such as the c-Met receptor, which has been implicated in the development and progression of several cancers. However, despite such progress, drug resistance continues to be the single most important cause of cancer treatment failure, and understanding the mechanisms of drug resistance remains a major hurdle in treating patients with recurrent disease. PF-04217903 is a small-molecule c-Met kinase inhibitor that potently inhibits c-Met-driven processes such as cell growth (proliferation and survival), motility, invasion, and morphology of a variety of tumor cells. Resistance to PF-04217903 was observed in GTL-16, a gastric carcinoma cell line with a constitutively activated c-Met receptor. In this report, mass spectrometry (MS) based quantitative phosphoproteomic analysis was used to determine changes in signaling pathways in the parental cells in response to c-Met inhibition and to investigate the changes in protein levels and related canonical pathways in both parental and PF-04217903 resistant (R3) clones in response to c-Met inhibition. The quantitative MS workflow included phosphoprotein enrichment of cell lysates from six treatment conditions: in-solution digestion, chemical labeling of peptides with a set of 6-plex isobaric tandem mass tags (TMT), HILIC fractionation, phosphopeptide enrichment, and nano LC-MS/MS on a LTQ-Orbitrap mass spectrometer. An investigation of these quantitative datasets using Ingenuity Pathways Analysis (IPA) revealed pathway changes in the various treatments that were consistent with previously observed transcriptomic and phenotypic changes. Proteomic analysis also revealed an increase in B-Raf expression in R3 clones. Expression profiling confirmed that B-Raf gene copy number was up-regulated and also indicated the presence of a mutated form of B-Raf. Using a bottom-up MS approach, SND-1 was identified as the B-Raf fusion partner. The discovery of this novel B-Raf fusion protein presents a novel target with potential clinical implications in the treatment of patients resistant to c-Met inhibitors.[4] PF-04217903 is a selective, ATP-competitive inhibitor of c-Met, designed to block HGF-induced c-Met activation and downstream signaling (PI3K-AKT, RAS-ERK pathways) in cancer cells [1] - In c-Met-amplified tumors, PF-04217903 showed greater in vitro and in vivo activity compared to c-Met-low tumors, supporting c-Met as a predictive biomarker for response [2][3] - Proteomic analysis revealed that PF-04217903 downregulates proteins involved in cell adhesion (e.g., E-cadherin) and angiogenesis (e.g., VEGF-A) in H441 cells, providing additional insights into its mechanism [4] |
Solubility Data
| Solubility (In Vitro) |
|
|||
| Solubility (In Vivo) |
Solubility in Formulation 1: 2 mg/mL (5.37 mM) in 10% DMSO + 40% PEG300 + 5% Tween80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), suspension solution; with sonication. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 20.0 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: 2 mg/mL (5.37 mM) in 10% DMSO + 90% (20% SBE-β-CD in Saline) (add these co-solvents sequentially from left to right, and one by one), suspension solution; with ultrasonication. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 20.0 mg/mL clear DMSO stock solution to 900 μL of 20% SBE-β-CD physiological saline solution and mix evenly. Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Solubility in Formulation 3: ≥ 2 mg/mL (5.37 mM) (saturation unknown) in 10% DMSO + 90% Corn Oil (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 20.0 mg/mL clear DMSO stock solution to 900 μL of corn oil and mix evenly. Solubility in Formulation 4: 1% DMSO+30% polyethylene glycol+1% Tween 80: 30 mg/mL (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 2.6854 mL | 13.4271 mL | 26.8543 mL | |
| 5 mM | 0.5371 mL | 2.6854 mL | 5.3709 mL | |
| 10 mM | 0.2685 mL | 1.3427 mL | 2.6854 mL |