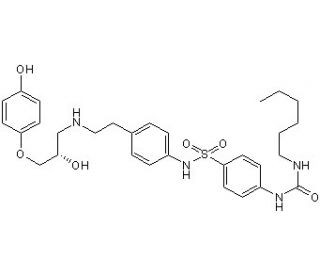
L-755,507 is identified as a potent and selective β3 adrenergic receptor partial agonist with EC50 of 0.43 nM, it displayed > 1000-fold selective over β1- and β2-adrenoceptors (EC50 values are 0.43, 580 and > 10000 nM for activation of cloned human β3-, β1- and β2-adrenoceptors respectively). It is also recently identified to enhance CRISPR-mediated homology-directed repair (HDR) efficiency in human induced pluripotent stem cells (iPSCs) and other cell types.
Physicochemical Properties
| Molecular Formula | C30H40N4O6S | |
| Molecular Weight | 584.73 | |
| Exact Mass | 584.266 | |
| CAS # | 159182-43-1 | |
| Related CAS # |
|
|
| PubChem CID | 9829836 | |
| Appearance | White to off-white solid powder | |
| Density | 1.3±0.1 g/cm3 | |
| Index of Refraction | 1.615 | |
| LogP | 4.97 | |
| Hydrogen Bond Donor Count | 6 | |
| Hydrogen Bond Acceptor Count | 8 | |
| Rotatable Bond Count | 17 | |
| Heavy Atom Count | 41 | |
| Complexity | 817 | |
| Defined Atom Stereocenter Count | 1 | |
| SMILES | CCCCCCNC(=O)NC1=CC=C(C=C1)S(=O)(=O)NC2=CC=C(C=C2)CCNC[C@@H](COC3=CC=C(C=C3)O)O |
|
| InChi Key | NYYJKMXNVNFOFQ-MHZLTWQESA-N | |
| InChi Code | InChI=1S/C30H40N4O6S/c1-2-3-4-5-19-32-30(37)33-24-10-16-29(17-11-24)41(38,39)34-25-8-6-23(7-9-25)18-20-31-21-27(36)22-40-28-14-12-26(35)13-15-28/h6-17,27,31,34-36H,2-5,18-22H2,1H3,(H2,32,33,37)/t27-/m0/s1 | |
| Chemical Name | 1-hexyl-3-[4-[[4-[2-[[(2S)-2-hydroxy-3-(4-hydroxyphenoxy)propyl]amino]ethyl]phenyl]sulfamoyl]phenyl]urea | |
| HS Tariff Code | 2934.99.9001 | |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
|
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
β3-Adrenoceptor (β3-AR) (EC50 = 1.8 nM for cAMP accumulation; EC50 = 3.2 nM for ERK1/2 phosphorylation) [1] |
| ln Vitro |
cAMP buildup increased significantly in response to L755507 (pEC50 values of 8.5 and 12.3, respectively). Maximal cAMP buildup rose following pretreatment with pertussis toxin and increased with zinterol and L755507. Extracellular signal-regulated kinase 1/2 (Erk1/2) is phosphorylated more strongly by zinterol, L755507, and L748337 than by cAMP (pEC50 values of 10.9, 11.7, and 11.6). Scr7 and L755507 did not considerably lower cell viability. Between 10 and 200 μM, Scr7 had no effect on the distribution of the cell cycle. L755507 significantly decreased the proportion of G2/M phase cells at 10 μM or 40 μM and increased the number of S phase cells at 10 μM in comparison to cells treated with DMSO [2]. In Chinese Hamster Ovary-K1 (CHO-K1) cells stably expressing human β3-AR, L755507 dose-dependently induces cAMP accumulation with an EC50 of 1.8 nM. At 100 nM, it achieves 89% of the maximal cAMP response induced by forskolin, and this effect is completely blocked by the β3-AR antagonist L748337 [1] - L755507 (0.1–100 nM) activates ERK1/2 MAPK signaling in CHO-K1-hβ3 cells, with an EC50 of 3.2 nM. At 10 nM, phosphorylated ERK1/2 (p-ERK1/2) levels are 2.7-fold higher than vehicle control, and the phosphorylation is attenuated by L748337 pretreatment [1] - In human primary T cells and mouse embryonic fibroblasts (MEFs), L755507 (1–10 μM) dose-dependently enhances CRISPR/Cas9-mediated homology-directed repair (HDR) efficiency. At 10 μM, HDR efficiency increases from 4.2% to 12.8% in human primary T cells and from 5.1% to 14.3% in MEFs, without significant impact on non-homologous end joining (NHEJ) activity [2] |
| ln Vivo |
Acute exposure of rhesus monkeys to L-755,507 elicits lipolysis and metabolic rate elevation, and that chronic exposure increases uncoupling protein 1 expression in rhesus brown adipose tissue. In medaka fish (Oryzias latipes) embryos, L755507 treatment significantly improves double gene knock-in efficiency. Embryos exposed to 5 μM L755507 for 24 hours post-fertilization (along with CRISPR/Cas9 components and donor DNA) show a double knock-in rate of 23.5%, compared to 8.7% in vehicle control. The embryo survival rate remains comparable (82% in treatment group vs 85% in control) [3] |
| Enzyme Assay |
cAMP accumulation assay: CHO-K1-hβ3 cells were seeded in 24-well plates and cultured for 24 hours, then serum-starved for 4 hours. Serial concentrations of L755507 (0.01–1000 nM) were added, and cells were incubated for 30 minutes. Cells were lysed with ice-cold lysis buffer, and intracellular cAMP levels were quantified using a competitive ELISA kit. EC50 values were calculated from dose-response curves of cAMP production [1] - ERK1/2 phosphorylation assay: CHO-K1-hβ3 cells were serum-starved for 16 hours, then treated with L755507 (0.1–100 nM) for 5 minutes. Cells were lysed, and proteins were separated by SDS-PAGE, transferred to PVDF membranes, and probed with anti-p-ERK1/2 and anti-total ERK1/2 antibodies. Band intensity was quantified by densitometry, and EC50 was derived from phosphorylation efficiency [1] |
| Cell Assay |
β3-AR signaling pathway assay: CHO-K1-hβ3 cells were plated in 6-well plates (2×105 cells/well) and cultured for 24 hours. After serum starvation, cells were pretreated with L748337 (1 μM) for 15 minutes (where applicable) followed by L755507 (0.1–100 nM) treatment. cAMP levels were measured by ELISA, and p-ERK1/2 expression was detected by western blot to confirm β3-AR-specific activation [1] - CRISPR/Cas9 HDR efficiency assay: Human primary T cells were isolated from peripheral blood and activated in culture for 3 days. Cells were electroporated with Cas9 plasmid, sgRNAs, and donor DNA, then immediately treated with L755507 (1–10 μM). After 72 hours of culture, genomic DNA was extracted, and the target region was amplified by PCR. HDR efficiency was determined by Sanger sequencing and analysis of mutation rates [2] |
| Animal Protocol |
Formulated in 25% ethanol, 25%polyethylene glycol 400, 50% saline; 3 mg/kg; i.v. injection Male lean rhesus monkeys Medaka fish double gene knock-in model: Mature medaka fish were allowed to mate naturally to obtain fertilized eggs. Embryos at 1–2 hours post-fertilization (hpf) were randomly divided into two groups (n=50/group): control and L755507 treatment. Treatment group embryos were immersed in embryo culture medium containing 5 μM L755507 for 24 hours. Meanwhile, all embryos were microinjected with Cas9 mRNA (200 ng/μL), two sgRNAs (100 ng/μL each), and donor DNA (50 ng/μL) into the cytoplasm. After 7 days of culture, 20 embryos per group were randomly selected, genomic DNA was extracted, and nested PCR and Sanger sequencing were performed to verify double gene knock-in efficiency. Embryo survival rates were recorded throughout the culture period [3] |
| References |
[1]. The beta3-adrenoceptor agonist 4-[[(Hexylamino)carbonyl]amino]-N-[4-[2-[[(2S)-2-hydroxy-3-(4-hydroxyphenoxy)propyl]amino]ethyl]-phenyl]-benzenesulfonamide (L755507) and antagonist (S)-N-[4-[2-[[3-[3-(acetamidomethyl)phenoxy]-2-hydroxypropyl]amino]-ethyl]phenyl]benzenesulfonamide (L748337) activate different signaling pathways in Chinese hamster ovary-K1 cells stably expressing the human beta3-adrenoceptor. Mol Pharmacol. 2008 Nov;74(5):1417-28. [2]. Small molecules enhance CRISPR/Cas9-mediated homology-directed genome editing in primary cells. Sci Rep. 2017; 7: 8943. [3]. An effective double gene knock-in strategy using small-molecule L755507 in the medaka fish (Oryzias latipes). Genesis. 2022;60(1-2):e23465. |
| Additional Infomation |
L755507 is a selective agonist of the β3-adrenoceptor (β3-AR), capable of activating distinct signaling pathways (cAMP/PKA and ERK1/2 MAPK) in cells expressing human β3-AR. Its activation profile differs from that of β3-AR antagonists, making it a valuable tool for studying β3-AR functional selectivity [1] - L755507 acts as a small-molecule enhancer of CRISPR/Cas9-mediated homology-directed repair (HDR) in primary cells, with low cytotoxicity. It improves HDR efficiency without affecting NHEJ, providing a practical tool for gene editing in primary cells [2] - In medaka fish embryos, L755507 effectively enhances double gene knock-in efficiency without reducing embryo survival, offering a feasible method for multi-gene editing in fish models [3] |
Solubility Data
| Solubility (In Vitro) |
|
|||
| Solubility (In Vivo) |
Solubility in Formulation 1: ≥ 2.5 mg/mL (4.28 mM) (saturation unknown) in 10% DMSO + 40% PEG300 + 5% Tween80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 25.0 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: ≥ 2.5 mg/mL (4.28 mM) (saturation unknown) in 10% DMSO + 90% (20% SBE-β-CD in Saline) (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 25.0 mg/mL clear DMSO stock solution to 900 μL of 20% SBE-β-CD physiological saline solution and mix evenly. Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Solubility in Formulation 3: ≥ 2.5 mg/mL (4.28 mM) (saturation unknown) in 10% DMSO + 90% Corn Oil (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 25.0 mg/mL clear DMSO stock solution to 900 μL of corn oil and mix evenly. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.7102 mL | 8.5510 mL | 17.1019 mL | |
| 5 mM | 0.3420 mL | 1.7102 mL | 3.4204 mL | |
| 10 mM | 0.1710 mL | 0.8551 mL | 1.7102 mL |