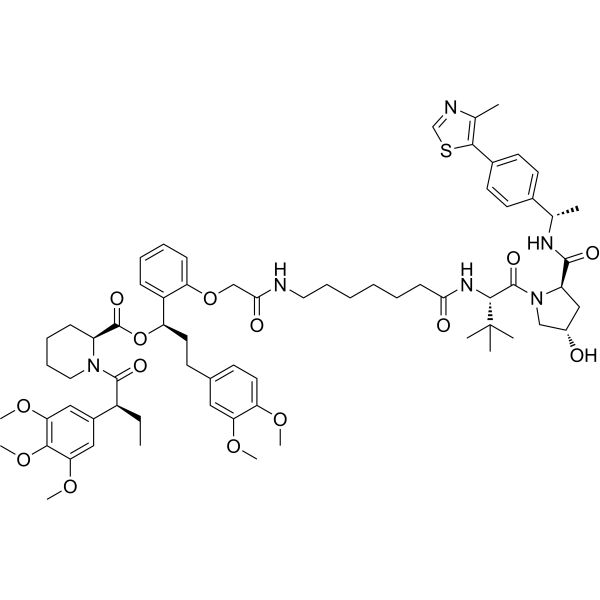
Physicochemical Properties
| Molecular Formula | C68H90N6O14S |
| Molecular Weight | 1247.53901815414 |
| Exact Mass | 1246.623 |
| Elemental Analysis | C, 65.47; H, 7.27; N, 6.74; O, 17.95; S, 2.57 |
| CAS # | 2451573-87-6 |
| Related CAS # | 2624313-15-9 (dTAGV-1 TFA); 2624313-16-0 (dTAGV-1 hydrochloride); 2451573-87-6 (dTAGV-1-NEG) |
| PubChem CID | 154828676 |
| Appearance | White to off-white solid powder |
| LogP | 10.1 |
| Hydrogen Bond Donor Count | 4 |
| Hydrogen Bond Acceptor Count | 16 |
| Rotatable Bond Count | 32 |
| Heavy Atom Count | 89 |
| Complexity | 2210 |
| Defined Atom Stereocenter Count | 7 |
| SMILES | CC[C@@H](C1=CC(=C(C(=C1)OC)OC)OC)C(=O)N2CCCC[C@H]2C(=O)O[C@H](CCC3=CC(=C(C=C3)OC)OC)C4=CC=CC=C4OCC(=O)NCCCCCCC(=O)N[C@H](C(=O)N5C[C@H](C[C@@H]5C(=O)N[C@@H](C)C6=CC=C(C=C6)C7=C(N=CS7)C)O)C(C)(C)C |
| InChi Key | ANLKEOUWAHUESE-SVUPEXKHSA-N |
| InChi Code | InChI=1S/C68H90N6O14S/c1-12-49(47-36-57(84-9)61(86-11)58(37-47)85-10)65(79)73-34-20-18-22-51(73)67(81)88-54(31-25-44-26-32-55(82-7)56(35-44)83-8)50-21-16-17-23-53(50)87-40-60(77)69-33-19-14-13-15-24-59(76)72-63(68(4,5)6)66(80)74-39-48(75)38-52(74)64(78)71-42(2)45-27-29-46(30-28-45)62-43(3)70-41-89-62/h16-17,21,23,26-30,32,35-37,41-42,48-49,51-52,54,63,75H,12-15,18-20,22,24-25,31,33-34,38-40H2,1-11H3,(H,69,77)(H,71,78)(H,72,76)/t42-,48-,49-,51-,52+,54+,63+/m0/s1 |
| Chemical Name | [(1R)-3-(3,4-dimethoxyphenyl)-1-[2-[2-[[7-[[(2S)-1-[(2R,4S)-4-hydroxy-2-[[(1S)-1-[4-(4-methyl-1,3-thiazol-5-yl)phenyl]ethyl]carbamoyl]pyrrolidin-1-yl]-3,3-dimethyl-1-oxobutan-2-yl]amino]-7-oxoheptyl]amino]-2-oxoethoxy]phenyl]propyl] (2S)-1-[(2S)-2-(3,4,5-trimethoxyphenyl)butanoyl]piperidine-2-carboxylate |
| Synonyms | dTAGV-1-NEG; 2451573-87-6; [(1R)-3-(3,4-Dimethoxyphenyl)-1-[2-[2-[[7-[[(2S)-1-[(2R,4S)-4-hydroxy-2-[[(1S)-1-[4-(4-methyl-1,3-thiazol-5-yl)phenyl]ethyl]carbamoyl]pyrrolidin-1-yl]-3,3-dimethyl-1-oxobutan-2-yl]amino]-7-oxoheptyl]amino]-2-oxoethoxy]phenyl]propyl] (2S)-1-[(2S)-2-(3,4,5-trimethoxyphenyl)butanoyl]piperidine-2-carboxylate; SCHEMBL25887266; |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets | VHL; FKBP12F36V fusion protein |
| ln Vitro | FKBP12F36V-Nluc is not degraded on 293FT FKBP12F36V-Nluc or FKBP12WT-Nluc cells by dTAGV-1-NEG (1-10 µM; 24 hours) [1]. PATU-8902 FKBP12F36V-KRASG12V and KRAS−/− cells do not exhibit KRASG12V protein degradation in response to dTAGV-1-NEG (500 nM; 24 hours) [1]. |
| Cell Assay |
Western Blot Analysis[1] Cell Types: PATU-8902 FKBP12F36V-KRASG12V, KRAS−/− Cell Tested Concentrations: 500 nM Incubation Duration: 1-24 hrs (hours) Experimental Results: KRASG12V protein degradation was not induced. Cell Viability Assay[1] Cell Types: EWS502 FKBP12F36V-GFP cells Tested Concentrations: 1 μM Incubation Duration: 0, 2, 4, 6, 8 days Experimental Results: demonstrated relative growth of cells same as the control group, higher than dTAGV-1 treated group. |
| References |
[1]. Rapid and direct control of target protein levels with VHL-recruiting dTAG molecules. Nat Commun. 2020 Sep 18;11(1):4687. |
| Additional Infomation |
Chemical biology strategies for directly perturbing protein homeostasis including the degradation tag (dTAG) system provide temporal advantages over genetic approaches and improved selectivity over small molecule inhibitors. We describe dTAGV-1, an exclusively selective VHL-recruiting dTAG molecule, to rapidly degrade FKBP12F36V-tagged proteins. dTAGV-1 overcomes a limitation of previously reported CRBN-recruiting dTAG molecules to degrade recalcitrant oncogenes, supports combination degrader studies and facilitates investigations of protein function in cells and mice. [1] We report dTAGV-1, a potent and exclusively selective VHL-recruiting degrader of FKBP12F36V-tagged proteins. dTAGV-1 displays improved PK/PD properties and serves as an optimized tool for in vivo applications. Through evaluation of mutant KRAS degradation in PDAC models, we show that either CRBN or VHL can be co-opted to alleviate the aberrant signaling coordinated by this oncoprotein. By contrast, we observed contextual differences in the ability of these E3 ubiquitin ligase complexes to degrade EWS/FLI. This is consistent with our recent report demonstrating effective degradation of a core mediator subunit (MED14) with dTAGV-1 in HCT116 cells, a context in which CRBN-recruiting dTAG molecules were not effective. We observed that rapid MED14 degradation abrogated lineage-specifying transcriptional circuits. Together, our studies provide support for use of dTAGV-1 to overcome the current limitations of the dTAG system, enabling evaluation of the direct effects of fusion proteins recalcitrant to CRBN-recruiting dTAG molecules. [1] Employing dTAGV-1 to study EWS/FLI, we demonstrate that VHL-mediated degradation of EWS/FLI rapidly alters downstream target protein expression and leads to pronounced growth defects in Ewing sarcoma cells, providing a powerful model system to investigate immediate consequences of EWS/FLI loss. This data supports that targeting EWS/FLI for degradation with direct-acting heterobifunctional degraders or molecular glues may be a tractable strategy and identifies potential combination strategies with BET bromodomain degraders. Together, the suite of dTAG molecules and paired controls provided in this study will facilitate evaluation of the functional consequences of precise posttranslational protein removal for an expanded target pool. The dTAG system enables rapid modulation of protein abundance and serves as a versatile strategy to determine whether targeted degradation is a promising drug development approach for a given target in vitro and in vivo. [1] |
Solubility Data
| Solubility (In Vitro) | May dissolve in DMSO (in most cases), if not, try other solvents such as H2O, Ethanol, or DMF with a minute amount of products to avoid loss of samples |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 0.8016 mL | 4.0079 mL | 8.0158 mL | |
| 5 mM | 0.1603 mL | 0.8016 mL | 1.6032 mL | |
| 10 mM | 0.0802 mL | 0.4008 mL | 0.8016 mL |