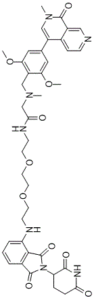
dBRD9 (dBRD-9) is a novel and potent PROTAC degrader of the BRD9 bromodomain and the cereblon E3 ubiquitin ligase complex with anticancer activity. The bromodomain-containing protein BRD9, a subunit of the human BAF (SWI/SNF) nucleosome remodeling complex, has emerged as an attractive therapeutic target in cancer. Despite the development of chemical probes targeting the BRD9 bromodomain, there is a limited understanding of BRD9 function beyond acetyl-lysine recognition. dBRD9, as the first BRD9-directed chemical degraders, was created through iterative design and testing of heterobifunctional ligands that bridge the BRD9 bromodomain and the cereblon E3 ubiquitin ligase complex. Degraders of BRD9 exhibit markedly enhanced potency compared to parental ligands (10- to 100-fold). Parallel study of degraders with divergent BRD9-binding chemotypes in models of acute myeloid leukemia resolves bromodomain polypharmacology in this emerging drug class. Together, these findings reveal the tractability of non-BET bromodomain containing proteins to chemical degradation, and highlight lead compound dBRD9 as a tool for the study of BRD9.
Physicochemical Properties
| Molecular Formula | C40H45N7O10 |
| Molecular Weight | 783.826209783554 |
| Exact Mass | 783.322 |
| CAS # | 2170679-45-3 |
| Related CAS # | dBRD9 dihydrochloride;2341840-98-8 |
| PubChem CID | 135397681 |
| Appearance | Light yellow to yellow solid powder |
| LogP | 0.9 |
| Hydrogen Bond Donor Count | 3 |
| Hydrogen Bond Acceptor Count | 13 |
| Rotatable Bond Count | 18 |
| Heavy Atom Count | 57 |
| Complexity | 1500 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | O=C1C(CCC(N1)=O)N1C(C2C=CC=C(C=2C1=O)NCCOCCOCCNC(CN(C)CC1C(=CC(=CC=1OC)C1=CN(C)C(C2C=NC=CC1=2)=O)OC)=O)=O |
| InChi Key | AIOCFZJGGGEWDK-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C40H45N7O10/c1-45(21-29-32(54-3)18-24(19-33(29)55-4)28-22-46(2)38(51)27-20-41-11-10-25(27)28)23-35(49)43-13-15-57-17-16-56-14-12-42-30-7-5-6-26-36(30)40(53)47(39(26)52)31-8-9-34(48)44-37(31)50/h5-7,10-11,18-20,22,31,42H,8-9,12-17,21,23H2,1-4H3,(H,43,49)(H,44,48,50) |
| Chemical Name | 2-((2,6-Dimethoxy-4-(2-methyl-1-oxo-1,2-dihydro-2,7-naphthyridin-4-yl)benzyl)(methyl)amino)-N-(2-(2-(2-((2-(2,6-dioxopiperidin-3-yl)-1,3-dioxoisoindolin-4-yl)amino)ethoxy)ethoxy)ethyl)acetamide |
| Synonyms | dBRD-9; dBRD 9; dBRD9 |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
dBRD9 targets BRD9 (bromodomain-containing protein 9, a subunit of the human BAF/SWI/SNF nucleosome remodeling complex) and cereblon (CRBN, a component of the E3 ubiquitin ligase complex). . [1] |
| ln Vitro |
Actin, BRD4, and BRD7 expression are not significantly affected by dBRD9 (0.5-5000 nM; 4 h), although it does concentrate the reduction of BRD9 expression in MOLM-13 cells [1]. In MOLM-13 cells, dBRD9 (100 nM; 2 h) is selective for BRD9 [1]. EOL-1 and MOML-13 cell lines exhibit anti-proliferative effects from dBRD9 (7 d) [1]. 1. dBRD9 is a heterobifunctional ligand that mediates the degradation of BRD9 through bridging BRD9 and the CRBN-DDB1 E3 ubiquitin ligase complex. It exhibits markedly enhanced potency (10- to 100-fold) compared to parental BRD9-targeting ligands. [1] 2. In MOLM-13 acute myeloid leukemia (AML) cells, dBRD9 induces dose-dependent degradation of BRD9 after 4 hours of treatment, as detected by immunoblotting (actin as a loading control). [1] 3. dBRD9 shows high biochemical and cellular selectivity: it does not significantly affect the expression of BRD7 or BRD4 (other bromodomain-containing proteins) in MOLM-13 cells, as confirmed by immunoblotting. Bromoscan analysis (phage-displayed bromodomain displacement assay) demonstrates its selectivity for BRD9 among bromodomains. [1] 4. Mechanistic studies in MM.1S cells show that dBRD9-induced BRD9 degradation is dependent on the ubiquitin-proteasome pathway: pre-treatment with the proteasome inhibitor carfilzomib, the NEDD8-activating enzyme inhibitor MLN-4924, or the CRBN ligand lenalidomide abrogates BRD9 degradation; pre-treatment with the BRD9 bromodomain inhibitor I-BRD9 also blocks this effect. Additionally, dBRD9 fails to degrade BRD9 in CRBN-knockout (MM.1SCRBN−/−) cells, confirming CRBN dependence. [1] 5. dBRD9 induces time-dependent BRD9 degradation in MOLM-13 cells: significant BRD9 reduction is observed after treatment with 100 nM dBRD9 for indicated time points (detected by immunoblotting). [1] 6. Whole-cell lysate proteomics analysis (quintuplicate replicates) of MOLM-13 cells treated with 100 nM dBRD9 for 2 hours (versus DMSO vehicle) confirms its selectivity: only BRD9 shows a significant fold change in relative abundance, with no major alterations in 7325 other quantified proteins. [1] 7. dBRD9 affects the viability of cultured human leukemia cell lines: treatment of EOL-1 and MOLM-13 cells with dBRD9 for 7 days reduces cell viability, as measured by ATP-Lite assay (quadruplicate means ± SEM). This effect is rescued by transduction of recombinant BRD9 alleles in MOLM-13 AML cells (vector control as reference). [1] 8. dBRD9 promotes the formation of a ternary complex between recombinant BRD9 bromodomain (BRD9(bd)) and CRBN-DDB1, as detected by AlphaScreen assay (quadruplicate means ± SEM). [1] |
| Enzyme Assay |
1. BRD9 bromodomain displacement assay (AlphaScreen): Recombinant BRD9 bromodomain (BRD9(bd)) is used to evaluate the displacement activity of dBRD9. The assay is performed in quadruplicate, and results are presented as vehicle-normalized displacement values with ± SEM. This assay measures the binding of dBRD9 to BRD9(bd). [1] 2. CRBN-DDB1 ternary complex formation assay (AlphaScreen): Recombinant BRD9(bd) and CRBN-DDB1 proteins are incubated with dBRD9, and the formation of the ternary complex is detected by AlphaScreen. The assay is conducted in quadruplicate, with results expressed as vehicle-normalized values ± SEM. A parallel assay with recombinant BRD4(1) bromodomain is used to confirm selectivity (no significant ternary complex formation with BRD4(1)). [1] 3. Bromodomain selectivity assay (Bromoscan): Phage-displayed bromodomains are used to assess the selectivity of dBRD9. The assay measures the displacement of dBRD9 from various bromodomains to confirm its preferential binding to BRD9. [1] |
| Cell Assay |
Western Blot Analysis[1] Cell Types: MOLM-13 cells. Tested Concentrations: 0.5, 5, 50, 500 and 5000 nM. Incubation Duration: 4 hrs (hours). Experimental Results: The expression of BRD9 was diminished. 1. BRD9 degradation detection by immunoblotting: MOLM-13 or MM.1S cells are treated with dBRD9 at indicated concentrations or time points (100 nM for time-course experiments). Cell lysates are prepared, and BRD9 protein levels are analyzed by immunoblotting, with actin as a loading control. For selectivity testing, BRD7 and BRD4 protein levels are also detected by immunoblotting in MOLM-13 cells. [1] 2. Mechanistic validation assay: MM.1S cells are pre-treated with vehicle, I-BRD9 (BRD9 inhibitor), lenalidomide (CRBN ligand), carfilzomib (proteasome inhibitor, 30-minute pre-treatment), or MLN-4924 (NEDD8-activating enzyme inhibitor) for 4 hours, followed by 2 hours of treatment with 100 nM dBRD9. BRD9 degradation is evaluated by immunoblotting (actin as control). Additionally, MM.1Swt and MM.1SCRBN−/− cells are treated with dBRD9 at indicated doses for 4 hours, and BRD9 levels are detected by immunoblotting. [1] 3. Cell viability assay: EOL-1 and MOLM-13 leukemia cells are treated with dBRD9 at indicated concentrations for 7 days. Cell viability is measured using the ATP-Lite assay, with results presented as quadruplicate means ± SEM. For rescue experiments, MOLM-13 cells are transduced with recombinant BRD9 alleles or vector control prior to dBRD9 treatment, and viability is assessed similarly. [1] 4. Whole-cell proteomics analysis: MOLM-13 cells are treated with 100 nM dBRD9 or DMSO vehicle for 2 hours (quintuplicate replicates). Whole-cell lysates are prepared, and 7326 proteins are quantified by proteomics. The fold change in relative abundance of each protein and corresponding q-values are calculated to assess the selectivity of dBRD9. [1] |
| References |
[1]. Degradation of the BAF Complex Factor BRD9 by Heterobifunctional Ligands. Angew Chem Int Ed Engl. 2017 May 15;56(21):5738-5743. |
| Additional Infomation |
1. dBRD9 is the first BRD9-directed chemical degrader, designed through iterative optimization of heterobifunctional ligands. [1] 2. The mode of action of dBRD9 involves recruiting the CRBN E3 ubiquitin ligase to BRD9, leading to BRD9 ubiquitination and subsequent degradation via the proteasome pathway. [1] 3. dBRD9 resolves bromodomain polypharmacology in BRD9-targeting agents and serves as a powerful tool for investigating BRD9 function beyond acetyl-lysine recognition. [1] 4. BRD9 has emerged as an attractive therapeutic target in cancer, particularly in acute myeloid leukemia. [1] 5. dBRD9 belongs to the class of naphthiridinone-based BRD9 degraders, with improved biochemical and cellular selectivity compared to earlier thienopyridinone-based degraders. [1] |
Solubility Data
| Solubility (In Vitro) | DMSO : ~100 mg/mL (~127.58 mM) |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.2758 mL | 6.3789 mL | 12.7579 mL | |
| 5 mM | 0.2552 mL | 1.2758 mL | 2.5516 mL | |
| 10 mM | 0.1276 mL | 0.6379 mL | 1.2758 mL |