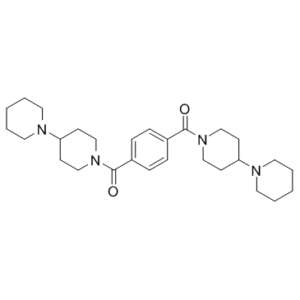
UNC1079, the piperidine analog of UNC1021, is a structurally similar but significantly less potent inhibitor for use as a negative control in cellular studies. It is used as a chemical probe for the methyllysine (Kme) reading function of L3MBTL3, a member of the malignant brain tumor (MBT) family of chromatin-interacting transcriptional repressors. The low anticipated affinity of UNC1079 was confirmed, as it demonstrated an activity versus L3MBTL3 of > 10 μM by AlphaScreen, which is >1000-fold weaker than UNC1215. UNC1079 also displays weak binding by ITC.
Physicochemical Properties
| Molecular Formula | C28H42N4O2 | |
| Molecular Weight | 466.66 | |
| Exact Mass | 466.33 | |
| Elemental Analysis | C, 72.07; H, 9.07; N, 12.01; O, 6.86 | |
| CAS # | 1418741-86-2 | |
| Related CAS # |
|
|
| PubChem CID | 70679311 | |
| Appearance | White to off-white solid powder | |
| Density | 1.2±0.1 g/cm3 | |
| Boiling Point | 640.0±55.0 °C at 760 mmHg | |
| Flash Point | 264.2±23.9 °C | |
| Vapour Pressure | 0.0±1.9 mmHg at 25°C | |
| Index of Refraction | 1.590 | |
| LogP | 2.28 | |
| Hydrogen Bond Donor Count | 0 | |
| Hydrogen Bond Acceptor Count | 4 | |
| Rotatable Bond Count | 4 | |
| Heavy Atom Count | 34 | |
| Complexity | 606 | |
| Defined Atom Stereocenter Count | 0 | |
| SMILES | C1(C(N2CCC(N3CCCCC3)CC2)=O)=CC=C(C(N2CCC(N3CCCCC3)CC2)=O)C=C1 |
|
| InChi Key | HOUMUCTZMMLNTR-UHFFFAOYSA-N | |
| InChi Code | InChI=1S/C28H42N4O2/c33-27(31-19-11-25(12-20-31)29-15-3-1-4-16-29)23-7-9-24(10-8-23)28(34)32-21-13-26(14-22-32)30-17-5-2-6-18-30/h7-10,25-26H,1-6,11-22H2 | |
| Chemical Name | [4-(4-piperidin-1-ylpiperidine-1-carbonyl)phenyl]-(4-piperidin-1-ylpiperidin-1-yl)methanone | |
| Synonyms |
|
|
| HS Tariff Code | 2934.99.03.00 | |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
|
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
L3MBTL3 methyllysine (Kme) reader domain [1] |
||
| ln Vitro |
In vitro activity: Kinase Assay: Cell Assay: 1. UNC1079 is a structurally similar analog of UNC1215 (a potent L3MBTL3 Kme reader domain antagonist) but exhibits significantly lower potency in antagonizing L3MBTL3 methyl-lysine binding activity; no specific inhibitory data (e.g., IC50, Kd) were provided [1] 2. Unlike UNC1215, UNC1079 does not competitively displace mono- or dimethyllysine-containing peptides from L3MBTL3 at effective concentrations of UNC1215 [1] 3. Isothermal titration calorimetry (ITC) analysis showed no significant binding of UNC1079 to wild-type L3MBTL3 (3MBT) protein, nor to domain mutants (D274A, D381A) of L3MBTL3 [1] |
||
| ln Vivo |
|
||
| Enzyme Assay |
1. ITC binding assay for L3MBTL3 and UNC1079: Wild-type L3MBTL3 (3MBT) protein and its domain mutants (D274A in domain 1, D381A in domain 2) were prepared; UNC1079 was titrated into the protein solution, and the heat change during the interaction was measured to evaluate binding affinity; no detectable binding of UNC1079 to L3MBTL3 (wild-type or mutants) was observed [1] 2. H4K20me2 pull-down assay with UNC1079: Biotinylated H4K20me2 peptides were immobilized, and L3MBTL3 (3MBT) protein was incubated with the peptides in the presence of gradient concentrations of UNC1079; the pulled-down L3MBTL3 was quantified, and UNC1079 showed no significant inhibition of L3MBTL3 binding to H4K20me2 (in contrast to UNC1215, which exhibited nanomolar potency) [1] |
||
| Cell Assay |
1. FRAP (Fluorescence Recovery After Photobleaching) assay in GFP-3MBT expressing HEK293 cells: Cells expressing GFP-tagged L3MBTL3 (3MBT) were treated with gradient concentrations of UNC1079; a specific nuclear area was photobleached, and the recovery time of fluorescence was recorded to evaluate the mobility of GFP-3MBTL3; UNC1079 showed no effect on the recovery time of photobleached GFP-3MBTL3 (whereas UNC1215 reduced the recovery time in a dose-dependent manner) [1] 2. Nuclear foci formation assay in HEK293 cells: HEK293 cells were transfected with GFP-3MBTL3 or GFP-FLMBTL3 (full-length L3MBTL3) plasmids, then treated with UNC1079 at different concentrations; the formation of nuclear foci of GFP-fusion proteins was observed under fluorescence microscopy; UNC1079 had no inhibitory effect on the foci formation of GFP-3MBTL3 (in contrast to UNC1215, which inhibited foci formation in a dose-dependent manner) [1] 3. Co-localization assay in U2OS cells: U2OS cells were transfected with GFP-3MBTL3 plasmid and treated with UNC1079; the co-localization of GFP-3MBTL3 and BCLAF1 was observed by immunofluorescence staining; UNC1079 did not disrupt the co-localization of 3MBTL3 and BCLAF1 (whereas UNC1215 disrupted the nuclear foci of 3MBTL3 and BCLAF1) [1] 4. Immunoprecipitation assay: HEK293 cells were transfected with flag-3MBTL3 or flag-FLMBTL3 plasmids, then treated with UNC1079; cell lysates were immunoprecipitated with anti-flag antibody, and the interaction between L3MBTL3 and BCLAF1 was detected by Western blot; UNC1079 did not disrupt the interaction between L3MBTL3 (3MBT/FLMBT) and BCLAF1 (whereas UNC1215 reduced this interaction) [1] |
||
| Animal Protocol |
|
||
| Toxicity/Toxicokinetics |
1. UNC1079 was non-toxic to HEK293 and U2OS cells at concentrations used in cellular experiments (no cell viability reduction or morphological changes were observed) [1] |
||
| References |
< a href="http://www.ncbi.nlm.nih.gov/pubmed/23292653">[1].Discovery of a chemical probe for the L3MBTL3 methyllysine reader domain. Nat Chem Biol. 2013 Mar;9(3):184-91. |
||
| Additional Infomation |
1. UNC1079 is a structural analog of UNC1215 (a potent and selective chemical probe for L3MBTL3 methyllysine reader domain) and serves as a negative control compound in L3MBTL3-related studies [1] 2. UNC1079 lacks the ability to bind to the Kme-binding pocket of L3MBTL3 MBT domains, which is the key functional region for UNC1215's antagonistic activity [1] 3. The structural similarity of UNC1079 to UNC1215 makes it an ideal negative control to verify the specific effects of UNC1215 on L3MBTL3-mediated biological processes (e.g., nuclear localization, protein-protein interaction with BCLAF1) [1] 4. UNC1079 does not interfere with the Kme-dependent interaction between L3MBTL3 and BCLAF1 (a protein involved in DNA damage repair and apoptosis) [1] |
Solubility Data
| Solubility (In Vitro) |
|
|||
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 2.1429 mL | 10.7144 mL | 21.4289 mL | |
| 5 mM | 0.4286 mL | 2.1429 mL | 4.2858 mL | |
| 10 mM | 0.2143 mL | 1.0714 mL | 2.1429 mL |