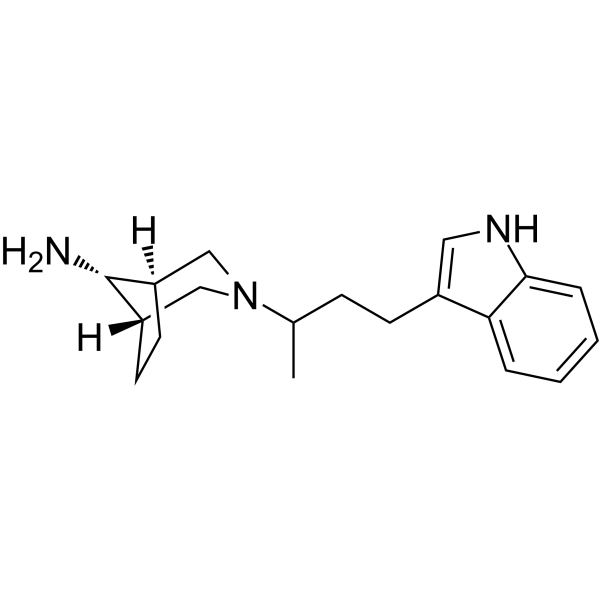
Physicochemical Properties
| Molecular Formula | C19H27N3 |
| Exact Mass | 297.22 |
| CAS # | 2103079-87-2 |
| Related CAS # | UHMCP1 dihydrochloride;2925647-93-2 |
| PubChem CID | 121513633 |
| Appearance | Typically exists as solid at room temperature |
| LogP | 3.2 |
| Hydrogen Bond Donor Count | 2 |
| Hydrogen Bond Acceptor Count | 2 |
| Rotatable Bond Count | 4 |
| Heavy Atom Count | 22 |
| Complexity | 371 |
| Defined Atom Stereocenter Count | 2 |
| SMILES | CC(CCC1=CNC2=CC=CC=C21)N3C[C@H]4CC[C@@H](C3)C4N |
| InChi Key | WCIFBVNRBHWVFY-XFROLERWSA-N |
| InChi Code | InChI=1S/C19H27N3/c1-13(22-11-15-8-9-16(12-22)19(15)20)6-7-14-10-21-18-5-3-2-4-17(14)18/h2-5,10,13,15-16,19,21H,6-9,11-12,20H2,1H3/t13?,15-,16+,19? |
| Chemical Name | (1S,5R)-3-[4-(1H-indol-3-yl)butan-2-yl]-3-azabicyclo[3.2.1]octan-8-amine |
| Synonyms | (1S,5R)-3-[4-(1H-indol-3-yl)butan-2-yl]-3-azabicyclo[3.2.1]octan-8-amine; UHMCP1; UHMCP-1 |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets | Kd: 79 µM (UHM domain)[1] |
| ln Vitro |
UHMCP1 (25-400 μM) exhibits a simple, non-cooperative binding mechanism with Kd of around 30 μM for U2AF65, reducing the binding of purified U2AF65 to SF3b155 by 75%[1]. UHMCP1 (0-200 μM; 24 h) has a specific influence on HEK293 cell viability[1]. UHMCP1 impacts splicing and cell viability Since U2AF is known to be essential for viability in many model organisms, and the UHM of U2AF65 is required for viability in S. pombe [[33]], we expected cell-permeant UHM-binding molecules to be toxic for cultured cells. We observed an effect of UHMCP1 on the viability of HEK293 cells as measured by a colorimetric MTS assay after 24 h of exposure to UHMCP1 (Fig. 8A). The toxicity was dependent on cell density, with an apparent EC50 of 140 µm with the highest density that was tested. We next analyzed the effect of UHMCP1 on splicing kinetics and alternative splicing by analyzing pre-mRNA accumulation and exon inclusion for a subset of genes. Total RNA was isolated after 16 h of exposure of HEK293 cells plated at high density, to 100 µm of UHMCP1. Cell morphology analysis by phase-contrast microscopy, just before RNA extraction, did not reveal any toxicity or differences between UHMCP1- and DMSO-treated control cells. This observation suggests that, if any splicing perturbations were detected, they were not due to an indirect effect of toxicity. Analysis of retention of randomly chosen introns in ACTB and GAPDH genes points to a decreased kinetics of splicing (Fig. 8B). In addition, several alternative splicing events were affected leading to increased or decreased skipping of some exons such as GALE exon 11 and CRYZ exon 7, respectively. In parallel, we also tested the effect of C29 that enhanced the U2AF65/SF3b155 interaction in cell extract. C29 had no effect on detection of the tested introns but could modulate exon inclusions differently from UHMCP1.[1] NMR analyses support the binding of UHMCP1 in the UHM hydrophobic pocket. WaterLOGSY and 15N-filtered NOESY experiments support the orientation of the UHMCP1 indole group in the UHM hydrophobic pocket. In silico experiments: molecular dynamics and free energy simulations support a stable orientation of UHMCP1 in the UHM hydrophobic pocket. [1] |
| Enzyme Assay |
NMR analyses [1] U2AF65 UHM spectra were recorded at 298 K on AVIII HD 600 Bruker NMR spectrometer, equipped with cryogenic triple resonance gradient probe. U2AF65 UHM backbone assignments were obtained from 2D 1H-15N-HSQC, 3D HNCO, 3D HN(CA)CO, 3D HNCA, 3D HN(CO)CA, 3D HNCACB, 3DHN(CO)CACB, 3D 1H-15N-NOESY-HSQC, and 3D 1H-15N-TOCSY-HSQC spectra and by comparison with previously reported NMR data. For NMR titration experiment, 2D 1H-15N SOFAST-HMQC spectra were recorded with 100 µm of U2AF65 UHM in presence of different UHMCP1 concentrations (25, 50, 100, 200, and 400 µm) in buffer H20N100. For each UHMCP1 concentration, chemical shift perturbations (CSPs) were calculated as urn:x-wiley:1742464X:media:febs16199:febs16199-math-0003. For Kd measurement, the averaged CSP values of the amino acids located inside the U2AF65 UHM hydrophobic pocket for different UHMCP1 concentrations were fitted using gosa-fit 3 software (Bio-Log, https://www.softpedia.com/get/Science-CAD/GOSA-fit.shtml). Assignment of UHMCP1 protons was performed using 2D HSQC and HMBC spectra on a 2 mm UHMCP1 sample in buffer H20N100 and P20N50. WaterLOGSY experiments were recorded using 1 mm of UHMCP1 in presence or absence of 20 µm U2AF65 UHM in buffer P20N50. WLOGSY factors were calculated by measuring changes in intensities upon protein binding as: WLOGSY factor = |(Iwlogsy+) – (Iwlogsy-)| / |(Iwlogsy-)|, with Iwlogsy+ and Iwlogsy- the peak intensities with and without protein. 2,2-Dimethyl-2-silapentane-5-sulfonic acid (DSS) was used for chemical shift referencing. Intermolecular NOEs between UHMCP1 and U2AF65 UHM were identified from a 2D F2 15N-filtered NOESY (τm 150 ms) experiment recorded using 250 µm 15N U2AF65 UHM and 460 µm UHMCP1 in buffer P20N50. Data were processed and analyzed with TOPSIN 4.0 (Bruker) and Ccpnmr Analysis V2 software. |
| Cell Assay |
Cell Viability Assay[1] Cell Types: HEK293 cells Tested Concentrations: 0-100 µM Incubation Duration: 24 h Experimental Results: Inhibited cell viability with an EC50 value of 140 µM. |
| References |
[1]. Identification of a small molecule splicing inhibitor targeting UHM domains. FEBS J. 2022 Feb;289(3):682-698. |
| Additional Infomation | Splicing factor mutations are frequent in myeloid neoplasms, blood cancers, and solid tumors. Cancer cells harboring these mutations present a particular vulnerability to drugs that target splicing factors such as SF3b155 or CAPERα. Still, the arsenal of chemical probes that target the spliceosome is very limited. U2AF homology motifs (UHMs) are common protein interaction domains among splicing factors. They present a hydrophobic pocket ideally suited to anchor small molecules with the aim to inhibit protein-protein interaction. Here, we combined a virtual screening of a small molecules database and an in vitro competition assay and identified a small molecule, we named UHMCP1 that prevents the SF3b155/U2AF65 interaction. NMR analyses and molecular dynamics simulations confirmed the binding of this molecule in the hydrophobic pocket of the U2AF65 UHM domain. We further provide evidence that UHMCP1 impacts RNA splicing and cell viability and is therefore an interesting novel compound targeting an UHM domain with potential anticancer properties.[1] |
Solubility Data
| Solubility (In Vitro) | May dissolve in DMSO (in most cases), if not, try other solvents such as H2O, Ethanol, or DMF with a minute amount of products to avoid loss of samples |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |