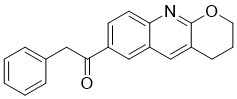
R214127 is a novel, potent and high-affinity radioligand for the mGlu1 receptor.
Physicochemical Properties
| Molecular Formula | C20H17NO2 |
| Molecular Weight | 303.36 |
| Exact Mass | 303.126 |
| Elemental Analysis | C, 79.19; H, 5.65; N, 4.62; O, 10.55 |
| CAS # | 409345-76-2 |
| Related CAS # | 409345-76-2; |
| PubChem CID | 10470232 |
| Appearance | Typically exists as solid at room temperature |
| LogP | 3.985 |
| Hydrogen Bond Donor Count | 0 |
| Hydrogen Bond Acceptor Count | 3 |
| Rotatable Bond Count | 3 |
| Heavy Atom Count | 23 |
| Complexity | 418 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | O1C2N=C3C(=CC=2CCC1)C=C(C(=O)CC1=CC=CC=C1)C=C3 |
| InChi Key | HXUSRWUBSYSWII-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C20H17NO2/c22-19(11-14-5-2-1-3-6-14)15-8-9-18-17(12-15)13-16-7-4-10-23-20(16)21-18/h1-3,5-6,8-9,12-13H,4,7,10-11H2 |
| Chemical Name | 1-(3,4-dihydro-2H-pyrano[2,3-b]quinolin-7-yl)-2-phenylethanone |
| Synonyms | R214127; R 214127; 409345-76-2; 1-(3,4-dihydro-2H-pyrano[2,3-b]quinolin-7-yl)-2-phenylethanone; R214,127; R 214,127; CHEMBL369459; Ethanone, 1-(3,4-dihydro-2H-pyrano[2,3-b]quinolin-7-yl)-2-phenyl-; [3H]R214127; [3H]-R214127; R-214127. |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets | mGlu1 |
| ln Vitro | R214127 was shown to be a potent and noncompetitive metabotropic glutamate 1 (mGlu1) receptor-selective antagonist. The kinetics and pharmacology of [(3)H]1-(3,4-dihydro-2H-pyrano[2,3-b]quinolin-7-yl)-2-phenyl-1-ethanone (R214127) binding to rat mGlu1a receptor Chinese hamster ovary (CHO)-dhfr(-) membranes was investigated, as well as the distribution of [(3)H]R214127 binding in rat brain tissue and sections. Specific binding to rat mGlu1a receptor CHO-dhfr(-) membranes was approximately 92% of total and was optimal at 4 degrees C. Full association was reached within 5 min, and [(3)H]R214127 bound to a single binding site with an apparent K(D) of 0.90 +/- 0.14 nM and a B(max) of 6512 +/- 1501 fmol/mg of protein. Inhibition experiments showed that [(3)H]R214127 binding was completely blocked by 2-quinoxaline-carboxamide-N-adamantan-1-yl (NPS 2390), (3aS,6aS)-6a-naphtalan-2-ylmethyl-5-methyliden-hexahydro-cyclopenta[c]furan-1-on (BAY 36-7620), and 7-(hydroxyimino)cyclo-propa[b]chromen-1a-carboxylate ethyl ester (CPCCOEt), but was not displaced by competitive mGlu1 receptor ligands such as glutamate and quisqualate, suggesting that R214127, NPS 2390, BAY 36-7620, and CPCCOEt bind to the same site or mutually exclusive sites. Experiments using rat cortex, striatum, hippocampus and cerebellum revealed that [(3)H]R214127 labeled a single high-affinity binding site (K(D) approximately 1 nM). B(max) values were highest in the cerebellum (4302 +/- 2042 fmol/mg of protein) and were 741 +/- 48, 688 +/- 125, and 471 +/- 68 fmol/mg of protein in the striatum, hippocampus, and cortex, respectively. The distribution of [(3)H]R214127 binding in rat brain was investigated in more detail by radioligand autoradiography. A high density of binding sites was detected in the molecular layer of the cerebellum. Moderate labeling was seen in the CA3 and dentate gyrus of the hippocampus, thalamus, olfactory tubercle, amygdala, and substantia nigra reticulata. The cerebral cortex, caudate putamen, ventral pallidum, and nucleus accumbens showed lower labeling. The high affinity and selectivity of [(3)H]R214127 for mGlu1 receptors renders this compound the ligand of choice to study the native mGlu1 receptor in brain. [2] |
| ln Vivo |
Metabotropic glutamate receptors mGluR5 and mGluR1 mediate key neuropsychiatric functions in health and disease and their antagonists hold promise to treat anxiety, depression, inflammation, and neuropathic pain. Pharmacological magnetic resonance imaging (phMRI) using a functional MRI approach in awake, conscious rodents can determine the activities of receptor ligands without the potential interference of anesthetics and independent of the specific biochemical mechanism of action of the candidate molecule. In this study we determined the neuronal activation patterns of 3-[(2-methyl-1,3-thiazol-4-yl)ethynyl]pyridine (MTEP) and 1-(3,4-dihydro-2H-pyrano[2,3-b]quinolin-7-yl0-2phenyl-1-ethanone (R214127), antagonists of mGluR5 and mGluR1 receptors by phMRI. We found that MTEP and R214127 activated specific primary somatosensory, piriform, entorhinal and motor cortices and the caudateputamen each to a different extent and in partly overlapping manners. Additional analysis of the activation data indicated that these brain regions and their connections are involved in mediating neuropathic pain and also, reward and olfaction. Using awake, conscious animals in phMRI can be a useful approach in characterizing candidate mGluR5 and mGlR1 antagonists also allowing a more direct comparison of animal and human phMRI studies. [1]
In contrast to the substantial activation of multiple brain regions caused by MTEP administration, R214127 treatment resulted in a significantly more restricted pattern and also a lower extent of overall activation (Fig. 1). The main brain regions activated by R214127 treatment are the piriform, entorhinal and motor cortices. In addition, there was activation in the pituitary. The activation pattern of R214127 we observed is only partly identical with previous data obtained by using classical methods. Using R214127 as a radioligand, a high density of binding was detected in the cerebellum (molecular layer), moderate in the CA3 and dentate gyrus of the hippocampus, thalamus, olfactory tubercle, amygdala and low in the cerebral cortex, caudate putamen. We excluded the cerebellum from the study because of the high incidence of motion artifacts due to respiration and swallowing that are present even in anesthesized animals. We saw no evidence of thalamic activation despite the high density of mGlR1 receptors described earlier. There can be several potential explanations for the observed discrepancy including, the specificity of the experimental ligand (R214127), the presence of endogenous mGluR1 ligand(s) and the known complexities of the positive BOLD signal. Earlier studies have shown the involvement of mGluR1 receptors in mediating long-term memory, cognition and pain perception. For detailed list of brain regions, their neuronal networks and known functions affected by R214127 (see Supplementary Table 1). Functional analysis of the primary topographic data using a segmented rat brain functional atlas indicated that the brain regions activated by MTEP and R214127 are primarily involved in mediating olfaction, reward and pain (Fig. 2). Further analysis regarding the involvement of the brain regions and their afferent and efferent pathways activated by MTEP and R214127 treatment indicated they are involved in wide ranges of neuronal functions (Supplementary Table 1). They included mediating various components of nociception processing of sensory, emotional and memory-related information of noxious stimuli. Our analysis has also illustrated some potentially important differences in the predicted functional consequences of MTEP and R214127 treatments. Accordingly, MTEP appears to affect more pain and pain-related information processing pathways than R214127 (Supplementary Fig. 2). These findings are consistent with previous data showing different distribution of mGluR1 and mGluR5 receptors and also their different involvements in pain and in other physiological and pathological processes [1]. |
| Enzyme Assay |
Selectivity and Mode of Antagonism of R214127 for the mGlu1 Receptor. [2] In CHO-dhfr− cells expressing the rat mGlu1a receptor, R214127 inhibited the glutamate-induced increase in [Ca2+]i with an IC50 value of 21.6 ± 5.0 nM (n = 4; Fig. 2A) and seemed to be about 8-fold more potent than the recently described specific mGlu1 receptor antagonist BAY 36-7620 (IC50 = 161 ± 38 nM, n = 3) and 500-fold more potent than CPCCOEt (IC50 = 10.3 ± 0.8 μM, n = 3), tested in the same assay. For the human mGlu1a receptor, R214127 had an IC50 value of 10.4 ± 4.7 nM (n = 3). |
| Animal Protocol | Adult male Long-Evan rats were housed in Plexiglas cages (two per cage) and maintained in ambient temperature (22–24 °C) on a 12:12 light:dark cycle (lights on at 09:00 h). Food and water were provided ad libitum. Briefly, animals were lightly anesthetized with isoflurane for securing the animal into the restrainer. When the animal regained full consciousness, the restraining unit was placed into a “mock scanner” and an audio tape recording of a typical MRI pulse sequence was played for 60 min per day. The noise intensity inside the mock scanner was similar to that of inside the magnet. The acclimation period lasted one week. All studies were performed using acclimated animals. Just prior to the imaging session, animals were briefly anesthetized with 2–3% isoflurane. A topical anesthetic of 10% lidocaine gel was applied to the skin and soft tissue around the ear canals and over the bridge of the nose. Studies were performed with a multi-concentric, dual-coil, small animal restrainer specifically developed for imaging fully awake, conscious rodents. R214127 (batch #: BZS 808) and MTEP were prepared as suspensions according to the manufacturer's instructions. The final concentrations used were 10 mg/kg of MTEP and 4 mg/kg of R214127. The doses were selected based on previous published reports and the drug manufacturer's in vivo internal studies (Richter Plc. Budapest, Hungary). The total volume of the i.p. injections were 500 μl and they were delivered over one minute. Rats (12 per treatment group) were randomly assigned to the following experimental conditions: (1) MTEP 10 mg/kg; (2) R214127 4 mg/kg; and (3) control. Animals in the control group received only the vehicle (5% Tween in PBS). Each animal was treated only once and with a single dose of the given drug or vehicle. At the end of the scan, the animal was euthanized using an overdose of the anesthetics followed by cervical dislocation. No animal was “reused”. The solutions containing the drugs were pretested for causing discomfort and/or pain prior to collecting imaging data. |
| References |
[1]. MRI analysis of mGluR5 and mGluR1 antagonists, MTEP and R214127 in the cerebral forebrain of awake, conscious rats. Neurosci Lett. 2011 Nov 14;505(2):155-9. [2]. [3H]R214127: a novel high-affinity radioligand for the mGlu1 receptor reveals a common binding site shared by multiple allosteric antagonists. Mol Pharmacol. 2003 May;63(5):1082-93. |
| Additional Infomation |
In conclusion, this is the first report that identifies the neuronal activation patterns following treatments with MTEP and R214127 in awake, conscious rats using phMRI. This is important because such approach reduces potential imaging artifacts caused by the use of anesthesia, it eliminates the known interference between anesthetics and glutamate-mediated synaptic transmission and enables a more direct comparison between animal and human clinical studies. In addition, such phMRI analysis can be completed within a few weeks. Activation data obtained following MTEP and R214127 treatments can be used as a “reference” for developing novel mGluR5 and mGluR1 antagonists. [1] Up to now, only a few mGlu1 receptor subtype-selective antagonists have been found. The mGlu1 receptor has been shown to be selectively blocked by CPCCOEt (Litschig et al., 1999) and BAY 36-7620 with potencies that vary from micromolar for CPCCOEt (6.6 μM) to high nanomolar for BAY 36-7620 (160 nM). In the present study, R214127 is identified as a novel mGlu1 receptor antagonist with low nanomolar functional antagonistic potency on the rat mGlu1a receptor (21.6 nM) and the human mGlu1a receptor [1]. |
Solubility Data
| Solubility (In Vitro) | May dissolve in DMSO (in most cases), if not, try other solvents such as H2O, Ethanol, or DMF with a minute amount of products to avoid loss of samples |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 3.2964 mL | 16.4821 mL | 32.9641 mL | |
| 5 mM | 0.6593 mL | 3.2964 mL | 6.5928 mL | |
| 10 mM | 0.3296 mL | 1.6482 mL | 3.2964 mL |