QNZ46 is a novel non-competitive and NR2C/NR2D-selective NMDA receptor antagonist with IC50 values of 3, 6, 229, and >300, >300 μM for NR2D, NR2C, NR2A, NR2B, and GluR1, respectively. NMDA receptors are ionotropic glutamate receptors that mediate excitatory synaptic transmission and have been implicated in several neurological diseases. QNZ46 inhibits NMDA receptor function in a noncompetitive and voltage-independent manner by an unconventional mechanism that requires binding of glutamate to the GluN2 subunit, but not glycine binding to the GluN1 subunit. Evaluation of the structural determinants responsible for the subunit-selectivity of QNZ46 revealed that these compounds act at a new site that has not previously been described. Residues residing in the part of the agonist binding domain immediately adjacent to the transmembrane helices appear to control selectivity of QNZ46 for GluN2C- and GluN2D-containing receptors. These residues are well-positioned to sense glutamate binding to GluN2 and thus to mediate glutamate-dependent actions.
Physicochemical Properties
| Molecular Formula | C24H17N3O6 | |
| Molecular Weight | 443.41 | |
| Exact Mass | 443.112 | |
| CAS # | 1237744-13-6 | |
| Related CAS # |
|
|
| PubChem CID | 46861929 | |
| Appearance | White to off-white solid powder | |
| LogP | 4.694 | |
| Hydrogen Bond Donor Count | 1 | |
| Hydrogen Bond Acceptor Count | 7 | |
| Rotatable Bond Count | 5 | |
| Heavy Atom Count | 33 | |
| Complexity | 814 | |
| Defined Atom Stereocenter Count | 0 | |
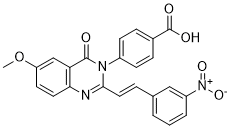
| SMILES | COC1=CC2=C(C=C1)N=C(N(C2=O)C3=CC=C(C=C3)C(=O)O)/C=C/C4=CC(=CC=C4)[N+](=O)[O-] |
|
| InChi Key | GNLVJIICVWDSNI-LFYBBSHMSA-N | |
| InChi Code | InChI=1S/C24H17N3O6/c1-33-19-10-11-21-20(14-19)23(28)26(17-8-6-16(7-9-17)24(29)30)22(25-21)12-5-15-3-2-4-18(13-15)27(31)32/h2-14H,1H3,(H,29,30)/b12-5+ | |
| Chemical Name | 4-[6-methoxy-2-[(E)-2-(3-nitrophenyl)ethenyl]-4-oxoquinazolin-3-yl]benzoic acid | |
| Synonyms |
|
|
| HS Tariff Code | 2934.99.9001 | |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
|
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
QNZ46 targets GluN2D-containing NMDA receptors (IC50 = 0.3 μM for recombinant rat GluN1/GluN2D receptors) [1] |
||
| ln Vitro |
In vitro activity: QNZ46 is a novel non-competitive and NR2C/NR2D-selective NMDA receptor antagonist with IC50 values of 3, 6, 229, and >300, >300 μM for NR2D, NR2C, NR2A, NR2B, and GluR1, respectively. NMDA receptors are ionotropic glutamate receptors that mediate excitatory synaptic transmission and have been implicated in several neurological diseases. QNZ46 inhibits NMDA receptor function in a noncompetitive and voltage-independent manner by an unconventional mechanism that requires binding of glutamate to the GluN2 subunit, but not glycine binding to the GluN1 subunit. Evaluation of the structural determinants responsible for the subunit-selectivity of QNZ46 revealed that these compounds act at a new site that has not previously been described. Residues residing in the part of the agonist binding domain immediately adjacent to the transmembrane helices appear to control selectivity of QNZ46 for GluN2C- and GluN2D-containing receptors. These residues are well-positioned to sense glutamate binding to GluN2 and thus to mediate glutamate-dependent actions. Kinase Assay: QNZ46 is a noncompetitive inhibitor of GluN2C/D containing NMDA receptors. KD and IC50 values for binding and inhibition of GluN1/Glun2D receptors by QNZ46 are 4.9 and 3.9 μM, respectively. QNZ46 does not compete for binding of glutamate or glycine, but QNZ46 receptor binding requires the binding of glutamate to the GluN2 subunit. Cell Assay: 1. QNZ46 potently inhibited glutamate/glycine-evoked currents in HEK293 cells expressing recombinant rat GluN1/GluN2D NMDA receptors with an IC50 of 0.3 μM; the inhibition was noncompetitive with respect to glutamate and glycine, indicating binding to an allosteric site [1] 2. QNZ46 showed minimal inhibition of GluN1/GluN2A, GluN1/GluN2B, and GluN1/GluN2C NMDA receptor subtypes at concentrations up to 10 μM, demonstrating high subtype selectivity for GluN2D-containing receptors [1] 3. Electrophysiological recordings revealed that QNZ46 reduced the peak amplitude of GluN1/GluN2D currents in a concentration-dependent manner (0.01-10 μM) and slowed current decay kinetics; the inhibitory effect was voltage-independent (no change in IC50 at holding potentials of -80 mV vs -40 mV) [1] 4. Mutagenesis studies identified key residues (Glu627, Ser631, Thr844) in the GluN2D subunit that are critical for QNZ46 binding and inhibition; mutation of these residues abolished or significantly reduced the inhibitory effect of QNZ46 on GluN1/GluN2D currents [1] 5. Molecular docking simulations showed that QNZ46 binds to a novel allosteric pocket in the GluN2D subunit, located at the interface between the ligand-binding domain (LBD) and the transmembrane domain (TMD) [1] |
||
| ln Vivo |
|
||
| Enzyme Assay |
1. Whole-cell patch-clamp electrophysiology assays were performed on HEK293 cells expressing recombinant rat NMDA receptor subtypes (GluN1/GluN2A, GluN1/GluN2B, GluN1/GluN2C, GluN1/GluN2D); cells were voltage-clamped at -80 mV (or -40 mV for voltage-dependence experiments), and glutamate/glycine (100 μM/10 μM) were applied to evoke currents; QNZ46 was pre-incubated with cells for 5 min at concentrations ranging from 0.01 μM to 10 μM, and peak current amplitude was measured to generate concentration-response curves and calculate IC50 values [1] 2. Site-directed mutagenesis was performed on the GluN2D subunit to generate mutant receptors (Glu627Ala, Ser631Ala, Thr844Ala); HEK293 cells were transfected with wild-type or mutant GluN1/GluN2D plasmids, and electrophysiological recordings were conducted to compare the inhibitory effect of QNZ46 on wild-type vs mutant receptors [1] 3. Molecular docking simulations were conducted to predict the binding mode of QNZ46 to the GluN2D subunit; the crystal structure of the GluN2D LBD-TMD interface was used as the template, and QNZ46 was docked into potential allosteric pockets using molecular modeling software; binding energy and residue interactions were analyzed to validate the binding site [1] |
||
| Cell Assay |
1. HEK293 cells were cultured in standard medium and transfected with plasmids encoding rat GluN1 and GluN2D (or other GluN2 subtypes) subunits; 24-48 h post-transfection, cells were plated on glass coverslips and used for electrophysiological recordings; cell viability was assessed before experiments to ensure >90% viability [1] 2. For kinetic analysis of GluN1/GluN2D currents, QNZ46 (0.1, 0.3, 1 μM) was applied to HEK293 cells expressing GluN1/GluN2D, and current decay time constants were calculated by fitting the current trace to a double-exponential function [1] 3. Mutant GluN2D subunits (Glu627Ala, Ser631Ala, Thr844Ala) were generated by site-directed mutagenesis; HEK293 cells were transfected with wild-type or mutant GluN1/GluN2D plasmids, and the expression of functional receptors was confirmed by measuring glutamate/glycine-evoked currents before testing the inhibitory effect of QNZ46 [1] |
||
| Animal Protocol |
|
||
| References |
[1]. Structural and mechanistic determinants of a novel site for noncompetitive inhibition of GluN2D-containing NMDA receptors. J Neurosci. 2011 Mar 9;31(10):3650-3661. |
||
| Additional Infomation |
1. NMDA receptors are ionotropic glutamate receptors composed of GluN1 and GluN2 (A-D) subunits; GluN2D-containing NMDA receptors are predominantly expressed in the developing brain and specific adult brain regions (e.g., midbrain, hindbrain), and are implicated in synaptic plasticity and neurological disorders [1] 2. QNZ46 is a novel noncompetitive allosteric inhibitor of GluN2D-containing NMDA receptors, binding to a unique site at the LBD-TMD interface of the GluN2D subunit [1] 3. The subtype selectivity of QNZ46 for GluN2D over other GluN2 subtypes is mediated by conserved residues in the GluN2D subunit that are not present in GluN2A/B/C; this selectivity makes QNZ46 a valuable tool compound for studying GluN2D receptor function [1] 4. Unlike competitive NMDA receptor antagonists (e.g., AP5), QNZ46 does not compete with glutamate or glycine binding, allowing it to modulate GluN2D activity without disrupting basal neurotransmission at other NMDA receptor subtypes [1] |
Solubility Data
| Solubility (In Vitro) |
|
|||
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 2.2552 mL | 11.2762 mL | 22.5525 mL | |
| 5 mM | 0.4510 mL | 2.2552 mL | 4.5105 mL | |
| 10 mM | 0.2255 mL | 1.1276 mL | 2.2552 mL |