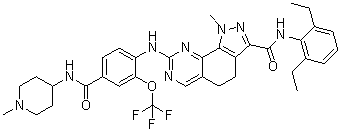
NMS-P715 (NMSP715) is a selective and orally bioactive MPS1 inhibitor with antitumor activity. It selectively reduces cancer cell proliferation, leaving normal cells almost unaffected. NMS-P715 accelerates mitosis and affects kinetochore components localization causing massive aneuploidy and cell death in a variety of tumorally bioavailable cell lines and inhibits tumor growth in preclinical cancer models. Inhibiting the SAC could represent a promising new approach to selectively target cancer cells.
Physicochemical Properties
| Molecular Formula | C35H39F3N8O3 |
| Molecular Weight | 676.73 |
| Exact Mass | 676.31 |
| Elemental Analysis | C, 62.12; H, 5.81; F, 8.42; N, 16.56; O, 7.09 |
| CAS # | 1202055-32-0 |
| Related CAS # | 1202055-34-2; 1202055-32-0 |
| PubChem CID | 44556162 |
| Appearance | Typically exists as White to off-white solids at room temperature |
| LogP | 6.293 |
| Hydrogen Bond Donor Count | 3 |
| Hydrogen Bond Acceptor Count | 11 |
| Rotatable Bond Count | 9 |
| Heavy Atom Count | 49 |
| Complexity | 1110 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | FC(OC1=C(C([H])=C([H])C(=C1[H])C(N([H])C1([H])C([H])([H])C([H])([H])N(C([H])([H])[H])C([H])([H])C1([H])[H])=O)N([H])C1=NC([H])=C2C(C3=C(C(C(N([H])C4C(=C([H])C([H])=C([H])C=4C([H])([H])C([H])([H])[H])C([H])([H])C([H])([H])[H])=O)=NN3C([H])([H])[H])C([H])([H])C2([H])[H])=N1)(F)F |
| InChi Key | JFOAJUGFHDCBJJ-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C35H39F3N8O3/c1-5-20-8-7-9-21(6-2)28(20)42-33(48)30-25-12-10-23-19-39-34(43-29(23)31(25)46(4)44-30)41-26-13-11-22(18-27(26)49-35(36,37)38)32(47)40-24-14-16-45(3)17-15-24/h7-9,11,13,18-19,24H,5-6,10,12,14-17H2,1-4H3,(H,40,47)(H,42,48)(H,39,41,43) |
| Chemical Name | N-(2,6-diethylphenyl)-1-methyl-8-[4-[(1-methylpiperidin-4-yl)carbamoyl]-2-(trifluoromethoxy)anilino]-4,5-dihydropyrazolo[4,3-h]quinazoline-3-carboxamide |
| Synonyms | NMSP715; NMSP715; NMSP715; NMS-P715; 1202055-32-0; N-(2,6-Diethylphenyl)-1-Methyl-8-({4-[(1-Methylpiperidin-4-Yl)carbamoyl]-2-(Trifluoromethoxy)phenyl}amino)-4,5-Dihydro-1h-Pyrazolo[4,3-H]quinazoline-3-Carboxamide; N-(2,6-diethylphenyl)-1-methyl-8-[4-[(1-methylpiperidin-4-yl)carbamoyl]-2-(trifluoromethoxy)anilino]-4,5-dihydropyrazolo[4,3-h]quinazoline-3-carboxamide; CHEMBL1236095; N-(2,6-diethylphenyl)-1-methyl-8-((4-((1-methylpiperidin-4-yl)carbamoyl)-2-(trifluoromethoxy)phenyl)amino)-4,5-dihydro-1H-pyrazolo[4,3-h]quinazoline-3-carboxamide; N-(2,6-Diethylphenyl)-1-methyl-8-[[4-[(1-methylpiperidin-4-yl)carbamoyl]-2-(trifluoromethoxy)phenyl]amino]-4,5-dihydro-1H-pyrazolo[4,3-h]quinazoline-3-carboxamide; NMSP715; |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets | Mps1 (IC50 = 182 nM); CK2 (IC50 = 5.7 μM); MELK (IC50 = 6.01 μM); NEK6 (IC50 = 6.02 μM) |
| ln Vitro | NMS-P715 has an IC50 of 182 nM, making it a selective inhibitor of MPS1. With only three kinases (CK2, MELK, and NEK6) inhibited below 10 μM and no other kinases inhibited below an IC50 value of 5 μM, NMS-P715 is highly specific for MPS1. With an EC50 of 65 nM, NMS-P715 encourages large spindle assembly checkpoint (SAC) overriding. NMS-P715 (1 μM) produces aneuploidy, slows the growth of HCT116 cells, and accelerates mitosis in U2OS cells that overexpress YFP-α-tubulin. NMS-P715 (0.5, 1 μM) influences the ubiquitylation of CDC20 and the stability of the mitotic checkpoint complex (MCC)[1]. NMS-P715 (1 μM) causes pancreatic ductal adenocarcinoma (PDAC) cell lines to undergo apoptosis and skip the spindle assembly checkpoint. Additionally, NMS-P715 (0-25 μM) specifically prevents PDAC cell growth[2]. |
| ln Vivo | Underneath mice with subcutaneously implanted human tumor cell xenografts demonstrate good pharmacokinetic characteristics and an oral bioavailability of 37% for NMS-P715 (10 mg/kg). In an A2780 ovarian cancer xenograft model, NMS-P715 (90 mg/kg, po) is well tolerated and does not cause any overt toxicities or symptoms of body weight loss. The A375 melanoma xenograft model shows that NMS-P715 (100 mg/kg, po) reduces tumor growth by approximately 43%[1]. |
| Enzyme Assay |
In Vitro Kinase Assays. [1] Recombinant full length MPS1 (2-857) was expressed as Nterminal, GST-tagged fusion protein using a baculovirus expression system. After cell lysis, the protein was loaded on a GST affinity column and after extensive washing, GST was cleaved with PreScission protease and the recombinant protein was eluted from the column. To fully activate MPS1, the protein was subjected to auto-phosphorylation in the presence of 1 mM ATP at 25°C for 2 hours in kinase buffer (50 mM Hepes pH 7.5, 2.5 mM MgCl2, 1 mM MnCl2, 1 mM DTT, phosphatase inhibitors); ATP was then removed with a desalting column. The potency of the compound towards MPS1 and 60 additional kinases belonging to our internal Kinase Selectivity Screening (KSS) panel was determined using either a strong anion exchanger (Dowex 1-X8 resin, formate form) based assay or P81 Multiscreen plate, both based on the specific measurement of radioactive phosphotransfer to the substrate, as previously described. For each KSS enzyme, the absolute Km values for ATP and the specific substrate were initially determined and each assay was then run at optimized [ATP] (2·αKm) and [substrate] (5·Km) concentrations. These conditions enabled direct comparison of IC50 values of NMSP715 or SP600125 across the KSS panel for the evaluation of biochemical selectivity. MPS1 activity was measured using 5 nM of MPS1 recombinant protein in 50 mM HEPES pH 7.5, 2.5 mM MgCl2, 1 mM MnCl2, 1 mM DTT, 3 μM NaVO3, 2 mM βglycerophosphate, 0.2 mg/mL BSA, 200 μM P38-βtide substrate-peptide (KRQADEEMTGYVATRWYRAE) and 8 μM ATP with 1.5 nM 33P-γ-ATP. The assay was run in a robotized format, 10 serial 1:3 compound dilutions (from 30 μM to 1.5 nM) were tested and IC50 determined. For time dependency and Ki evaluation experiments, 1 nM of MPS1 was used in the assay in order to use a simplified equation. Preincubation was performed 30 minutes before reaction start and was then arrested after 60 minutes.[1] Surface Plasmon Resonance.[1] Experiments were performed with Biacore T100. MPS1 full length enzyme (40 μg/ml) was immobilized on Sensor Chip CM5 using amine coupling according to the following procedure: 1) activation with 0.2 M Nethyl-N-dimethylaminopropylcarbodiimide and 50 mM N-hydroxysuccinimide for 10 min; 2) injection of MPS1 (5 μl/min) diluted in 10 mM HEPES pH 7.0, 2.5 mM MgCl2, 1 mM MnCl2, 1 mM DTT in the presence of 1 μM Staurosporine, until a workable level of enzyme immobilization on surface was obtained, followed by blockage with 1.0 M ethanolamine (pH 8.5) for 10 minutes. The reference surface was generated using the same procedure with the exception of the ligand injection step. Binding experiments were performed as previously described. |
| Cell Assay |
Cell growth inhibition assays[2] The structurally defined inhibitor, NMS-P715, was suspended in DMSO. Gemcitabine was suspended in H2O. Drug dose-response assays were performed by plating 2,000 human or 1,000 KRC cells per well in triplicate in 96 well plates. Three replicate assays were performed per cell line. Compounds were added for 72h after which cells were methanol fixed and stained with 0.05% methylene blue. Optical density was measured at 620 nM after suspension in 0.5M HCl on a Beckman-Coulter DTX880 MultiMode Detector. Proliferation was measured relative to vehicle control and IC50 determined using Compusyn software. Dose-response curves were generated using sigmoidal interpolation curve fitting in SigmaPlot 12.3. For clonogenic survival assays, cells were plated at the indicated densities in duplicate or triplicate in 12-well dishes and allowed to attach for 24h. For continuous treatment, NMS-P715 was replenished every three days. In the washout assays, cells received a 24h NMS-P715 treatment followed by culture in compound-free medium. Experiments were performed in duplicate. Cell growth quantification in the colony formation assay was by colorimetric methylene blue assay or manual counting. Growth inhibition was measured relative to vehicle control. Cell proliferation assay. [1] Cells lines were seeded in 384 well-plates in the appropiate complete medium and treated with NMS-P715 dissolved in 0.1% DMSO 24 hours after seeding. The cells were incubated at 37°C and 5% CO2 and after 72 hours the plates were processed using CellTiter-Glo assay. |
| Animal Protocol |
Animal efficacy studies.[1] Athymic nu-nu mice, 5–6 weeks of age (20–22 g) were obtained from Harlan, Italy. A2780 ovary carcinoma and A375 melanoma cells were transplanted s.c. into female nu-nu mice. Mice bearing a palpable tumor (100–200 mm3 ) were selected and randomized into control and treated groups. Treatment started one day after randomization. NMS-P715 was typically administered by oral administration at doses of 90-100 mg/kg daily for more than seven days. Each group included 8 animals. Tumor dimension was measured regularly by calipers during the experiments and tumor mass was calculated as described: the tumor growth inhibition (TGI, %) was calculated according to the equation: % TGI = 100 – (mean tumor weight of treated group/mean tumor weight of control group) * 100. Pharmacokinetics. [1] The PK of NMS-P715 was investigated in an ancillary group of three mice. Blood samples were collected and drug concentration was measured in plasma with liquid chromatography tandem mass spectrometry as previously described. |
| References |
[1]. Targeting the mitotic checkpoint for cancer therapy with NMS-P715, an inhibitor of MPS1 kinase. Cancer Res. 2010 Dec 15;70(24):10255-64. [2]. Selective inhibition of pancreatic ductal adenocarcinoma cell growth by the mitotic MPS1 kinase inhibitor, NMS-P715.Mol Cancer Ther. 2014 Feb;13(2):307-315. |
| Additional Infomation |
MPS1 kinase is a key regulator of the spindle assembly checkpoint (SAC), a mitotic mechanism specifically required for proper chromosomal alignment and segregation. It has been found aberrantly overexpressed in a wide range of human tumors and is necessary for tumoral cell proliferation. Here we report the identification and characterization of NMS-P715, a selective and orally bioavailable MPS1 small-molecule inhibitor, which selectively reduces cancer cell proliferation, leaving normal cells almost unaffected. NMS-P715 accelerates mitosis and affects kinetochore components localization causing massive aneuploidy and cell death in a variety of tumoral cell lines and inhibits tumor growth in preclinical cancer models. Inhibiting the SAC could represent a promising new approach to selectively target cancer cells.[1] Most solid tumors, including pancreatic ductal adenocarcinoma (PDAC), exhibit structural and numerical chromosome instability (CIN). Although often implicated as a driver of tumor progression and drug resistance, CIN also reduces cell fitness and poses a vulnerability that can be exploited therapeutically. The spindle assembly checkpoint (SAC) ensures correct chromosome-microtubule attachment, thereby minimizing chromosome segregation errors. Many tumors exhibit upregulation of SAC components such as MPS1, which may help contain CIN within survivable limits. Prior studies showed that MPS1 inhibition with the small molecule NMS-P715 limits tumor growth in xenograft models. In cancer cell lines, NMS-P715 causes cell death associated with impaired SAC function and increased chromosome missegregation. Although normal cells appeared more resistant, effects on stem cells, which are the dose-limiting toxicity of most chemotherapeutics, were not examined. Elevated expression of 70 genes (CIN70), including MPS1, provides a surrogate measure of CIN and predicts poor patient survival in multiple tumor types. Our new findings show that the degree of CIN70 upregulation varies considerably among PDAC tumors, with higher CIN70 gene expression predictive of poor outcome. We identified a 25 gene subset (PDAC CIN25) whose overexpression was most strongly correlated with poor survival and included MPS1. In vitro, growth of human and murine PDAC cells is inhibited by NMS-P715 treatment, whereas adipose-derived human mesenchymal stem cells are relatively resistant and maintain chromosome stability upon exposure to NMS-P715. These studies suggest that NMS-P715 could have a favorable therapeutic index and warrant further investigation of MPS1 inhibition as a new PDAC treatment strategy.[2] |
Solubility Data
| Solubility (In Vitro) | May dissolve in DMSO (in most cases), if not, try other solvents such as H2O, Ethanol, or DMF with a minute amount of products to avoid loss of samples |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.4777 mL | 7.3885 mL | 14.7769 mL | |
| 5 mM | 0.2955 mL | 1.4777 mL | 2.9554 mL | |
| 10 mM | 0.1478 mL | 0.7388 mL | 1.4777 mL |