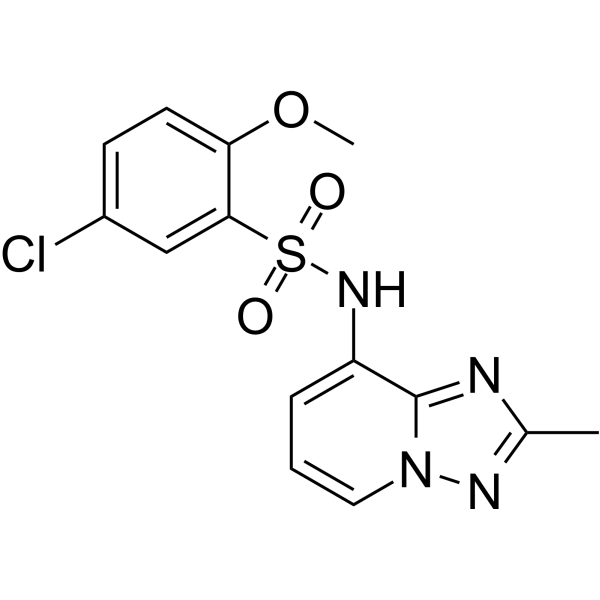
Physicochemical Properties
| Exact Mass | 352.04 |
| Elemental Analysis | C, 47.66; H, 3.71; Cl, 10.05; N, 15.88; O, 13.60; S, 9.09 |
| CAS # | 1341030-00-9 |
| PubChem CID | 87054166 |
| Appearance | Off-white to pink solid powder |
| LogP | 2.3 |
| Hydrogen Bond Donor Count | 1 |
| Hydrogen Bond Acceptor Count | 6 |
| Rotatable Bond Count | 4 |
| Heavy Atom Count | 23 |
| Complexity | 515 |
| Defined Atom Stereocenter Count | 0 |
| InChi Key | HLRMNXITZQAUIH-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C14H13ClN4O3S/c1-9-16-14-11(4-3-7-19(14)17-9)18-23(20,21)13-8-10(15)5-6-12(13)22-2/h3-8,18H,1-2H3 |
| Chemical Name | 5-chloro-2-methoxy-N-(2-methyl-[1,2,4]triazolo[1,5-a]pyridin-8-yl)benzenesulfonamide |
| Synonyms | C26-A6; C26 A6; MTDH-SND1 blocker 1; 5-chloro-2-methoxy-N-(2-methyl[1,2,4]triazolo[1,5-a]pyridin-8-yl)benzene-1-sulfonamide; EX-A8995; AKOS024452368; HY-155170; CS-0890400; 5-chloro-2-methoxy-N-(2-methyl[1,2,4]triazolo[1,5-a]pyridin-8-yl)-1-benzenesulfonamide; 1341030-00-9; |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets | MTDH-SND1 Protein[1] |
| ln Vitro |
C26s block SND1 pocket to disrupt MTDH-SND1 complex.[2] C26s inhibit breast cancer progression and metastasis.[2] To confirm at the molecular level that C26-A6 targets the MTDH-SND1 complex to suppress breast cancer progression, PyMT;UBC-CreERT+/−;Mtdhfl/fl mice with tumors were treated with vehicle, Tmx, and C26-A6, followed by next-generation RNA sequencing (NGS). Non-supervised clustering based on global gene expression data indicated that Tmx and C26-A6 groups cluster together (Extended Data Fig. 7j), suggesting that Mtdh acute KO and C26-A6 treatment have similar effect on gene expression in tumors. Moreover, gene set enrichment analysis (GSEA) showed that Tmx treatment upregulated and downregulated genes are sharply enriched positively or negatively in C26-A6 tumors, respectively (Fig. 6e, left and middle panels), suggesting Mtdh acute KO and C26-A6 treatment regulate similar genes and pathways in breast tumors. To provide a better understanding of how C26-A6 exerts its tumor suppressive function, C26-A6 downregulated genes in comparison with vehicle were employed for Ingenuity Pathway Analysis (IPA). A significant enrichment of “Cell Death and Survival”, “Cell Cycle”, and “DNA repair” molecular and cellular functions was observed (Extended Data Fig. 7k). Genes involved in cell survival and viability were decreased, whereas genes involved in apoptosis were increased (Extended Data Fig. 7l) in C26-A6 treated tumors, which is similar as what we found in SND1-dependent signature during chemotherapy (SND1_CPT_UP)4. Furthermore, SND1_CPT_UP signature that negatively enriched in MTDH-SND1 interaction deficient tumors4, is also down regulated upon C26-A6 treatment (Fig. 6e, right). Taken together, these results confirm that C26-A6 targets MTDH-SND1 interactions to exert a global gene expression changes, leading to inhibition of breast cancer progression.[2] |
| ln Vivo | To this end, Balb/C mice were injected with 4T1 mouse mammary tumor cells, which mimic TNBC33, via tail-vein and were subjected to the same treatment regime as above three days later. Consistently, C26-A6+paclitaxel significantly inhibited lung metastasis more than either treatment alone (Fig. 7g). Furthermore, mice with C26-A6+paclitaxel treatment had the best survival rate (Fig. 7h). Lastly, to mimic the clinical scenario of adjuvant chemotherapy following surgical removal of the primary tumors, SCP28 mammary tumors were removed when they reached 5 mm in diameter, the mice were then randomly separated into four groups and subjected to the different treatment regimens as above. Again, C26-A6+paclitaxel treatment achieved the more effective reduction of lung metastasis and overall survival (Fig. 7i,,j),j), suggesting targeting MTDH-SND1 complex together with chemotherapy could significantly improve the treatment outcome for breast cancer. To further evaluate if the treatment could result in the shrinkage of established macrometastases, we performed tail vein injections to generate 4TO7 lung metastases bearing mice. The mice were randomized into four groups of 6 mice each when macrometasetases were well-established, as evidenced by robust BLI signals in the lungs (Fig. 8a,,b).b). The mice were then treated with vehicle or paclitaxel and C26-A6 alone or in combination, and the metastasis progression was monitored by BLI. Although paclitaxel or C26-A6 treatment alone slowed down metastatic growth, these monotherapies did not result in metastasis shrinkage (Fig. 8c). However, three mice in C26-A6+paclitaxel group have stabilized disease and one mouse has obvious metastasis shrinkage, leading to significantly improved survival rate (Fig. 8c,,d).d). More importantly, consistent with our results showed above (Extended Data Fig. 6d–h), C26-A6 was well tolerated by the mice and did not further enhance the chemotherapy toxicity when combined with paclitaxel (Fig. 8e–h). These data suggest that C26-A6 combined with chemotherapy may have clinical benefit in metastatic breast cancer patients[2]. |
| Enzyme Assay |
FRET- based screening[2] To perform FRET assay, purified CFP-MTDH (386–407aa) and TC-SND1 (16–339aa) proteins were reconstituted in FRET buffer (25mM Tris-HCl pH8.0, 150 mM NaCl, 3mM DTT, 2% DMSO). 384-well plate was used for this assay and the compounds/peptides were distribute into each well as above. 0.065 μl of CFP-MTDH was added to 8.9755 μl of FRET buffer, and then transferred the mixture to each well. Incubate the plate for 5 min at room temperature, avoid light. 1.86 μl of TC-SND1, 0.024 μl of FIAsH-EDT2 Labeling reagent (TC-FlAsH™ II In-Cell Tetracysteine Tag Detection Kits, Cat#T34561) together with 8.9755 μl of FRET buffer was mixed and then added to each well. The plate was measured with excitation weave length of 450 nm and emission weavelength of 495 nm and 535 nm.[2] The inhibitory efficiency was calculated as following: After subtracting the value of DMSO background, the emission of CFP-MTDH at 495 nm is considered as D, the emission of TC-SND1+FIAsH is A, the emission of CFP-MTDH + TC-SND1+FIAsH mixture is DA, the efficiency is calculated as 1-(DA-A)/D. Similarly, MTDH wild type or mutant peptide in each plate was served as positive and negative controls to monitor the data quality of each plate.[2] Thermal Melt assay[2] 0.1 mg/ml of purified SND1 (16–339aa) protein together with 5-fold of Sypro Orange dye was added into assay buffer (20 mM HEPES, 150 Mm NaCl, pH 7.0). 250 μM of MTDH wild type/mutant peptides, indicated amount of compounds or same amount of DMSO was then added into each reaction. Temperature increased from 25°C to 95°C and fluorescence signal was measured. ΔTm max = Tm max (compound) - Tm max (DMSO)[2] Microsacle Thermophoresis (MST) assay[2] To perform MST assay, 150 nM purified SND1 (16–339aa) protein was labeled with 50 nM of RED-tris-NTA dye. Monolith NT.115 Instrument using the RED detector was employed. MTDH wild type/mutant peptides were screened in a twelve point two-fold serial dilution with concentrations ranging from 50 μM to 24.4 nM. The C26s compounds were screened in a twelve point three-fold serial dilution with concentrations ranging from 0.5 mM to 2.82 nM.[2] The assay was performed in buffer containing: 20 mM Hepes pH 7.0, 150 mM NaCl and 10 % Glycerol. After a 30 min incubation with the respective compound and 5 min centrifugation at 14000 rpm, the samples were loaded into MonolithTM NT.115 Series Standard Treated Capillaries. The MST analysis was performed using LED Power: 60% and MST Power: 40%. The normalized fluorescence Fnorm measures mainly this concentration ratio, plus a temperature dependence of the dye as noted before50. Concentrations on the x-axis are plotted in nM. A KD was determined with the MO.Affinity Analysis Software v2.2.4. |
| Cell Assay |
Luciferase-based screening[2] Seed HEK293T cells on 3×15 cm dish at 18–24 hr before transfection with 7–8X106 cells per dish targeting 70–80% confluence when start transfection. 20 μg of pCDNA3.1-SND1-NLuc, pCDNA3.1-CLuc-MTDH, or 2 μg of pCDNA3.1-NLuc-CLuc plasmids that described above were transfected to each dish. 72 hr after transfectionthe cells were lysed with 5 ml of luciferase lysis buffer (2 mM EDTA, 20 mM DTT, 10% glycerol, 1% Triton X-100, 25 mM Tris base, adjusted to pH7.8 with H3PO4) at 4°C for 20 min. The protein lysates were centrifuged at 13,000 rpm for 10 min, and the supernatant were collected.[2] For small chemical compound screening, white, flat-bottom, solid 384-well plates were used. Compounds were added into each well as 0.1 μl of 10 mM DMSO solution (or same amount of DMSO, serve as control). The first and last columns were free, and MTDH wild type (PNSDWNAPAEEWGNW) or mutant (PNSDANAPAEEAGNW) peptides were added as positive and negative controls right before screening. Same amount of MTDH-CLuc and SND1-NLuc were pre-mixed at 4°C for 30 min to generate split-luc. 5 μl of split- or linked-luc was added into each well. 15 μl of luciferase assay buffer (25 mM Glycylglycine pH 7.8, 15 mM K3PO4 pH 7.8, 15 mM MgSO4, 4 mM EGTA, 2 mM ATP added just before use, 10 mM DTT added just before use and 1 mM D-Luciferin added just before use) was then added into each well to get a 20 μl reaction system with 50 μM of compounds (peptides) in each well. The plates were incubated at 4°C for 1 hr and luciferase activity at each well was measured.[2] The inhibitory efficiency of each compound was calculated as following: (signal at DMSO well – signal at compound well) / signal at DMSO well. MTDH wild type or mutant peptide in each plate was served as positive and negative controls to monitor the data quality of each plate. |
| Animal Protocol | For C26-A6 treatments, mice were injected with 15 mg/kg via i.v. Primary tumors were considered established when they became palpable for 2 consecutive weeks. The tumors were measured by calipers for calculation of tumor volumes (length x width2/2). For cell lines that are stably labeled with a firefly luciferase expressing vector, lung metastases were monitored by bioluminescent imaging (BLI) and images were processed with Living Image 3D Analysis (version 1.0). For PDX treatment, fresh TNBC patient-derived xenograft (PDX, HCI-001) was chopped into ~1–2 mm in diameter cubes and inoculated into 8-week NSG females. One day after inoculation, the mice were randomized and treated with vehicle or C26-A6[2]. |
| References |
[1].Inhibitors of the mtdh-snd1 protein complex for cancer therapy. WO2023064669A3.2023-09-21. [2]. Small Molecule Inhibitors that Disrupt the MTDH-SND1 Complex Suppress Breast Cancer Progression and Metastasis. Nat Cancer. 2022 Jan; 3(1): 43–59. |
| Additional Infomation | Metastatic breast cancer is leading health burden worldwide. Previous studies have shown that Metadherin (MTDH) promotes breast cancer initiation, metastasis and therapy resistance; however, the therapeutic potential of targeting MTDH remains largely unexplored. Here, we used genetically modified mice and demonstrate that genetic ablation of Mtdh inhibits breast cancer development through disrupting the interaction with Staphylococcal nuclease domain-containing 1 (SND1) which is required to sustain breast cancer progression in established tumors. We performed a small molecule compound screening to identify a class of specific inhibitors that disrupt the protein-protein interaction between MTDH-SND1, and show that our lead candidate compounds C26-A2 and C26-A6 suppressed tumor growth and metastasis, and enhanced chemotherapy sensitivity in preclinical models of triple-negative breast cancer. Our results demonstrate a significant therapeutic potential in targeting the MTDH-SND1 complex and identify a new class of therapeutic agents for metastatic breast cancer.[2] |
Solubility Data
| Solubility (In Vitro) | Typically soluble in DMSO (e.g. 10 mM) |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |