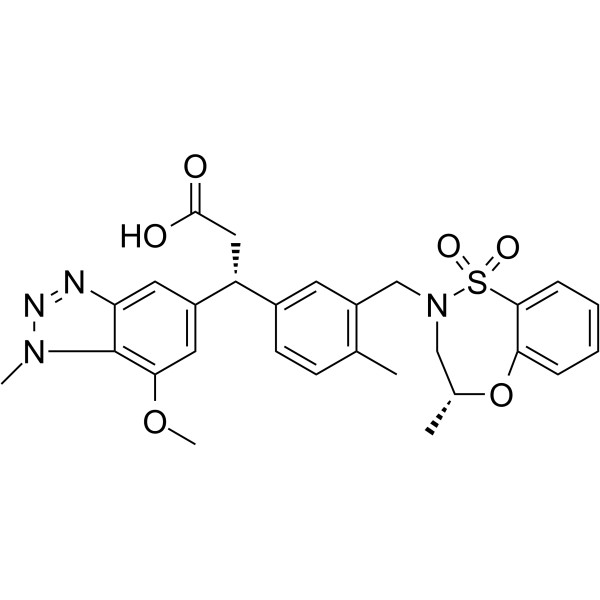
Physicochemical Properties
| Molecular Formula | C28H30N4O6S |
| Molecular Weight | 550.626005649567 |
| Exact Mass | 550.188 |
| CAS # | 1799974-69-8 |
| PubChem CID | 118169940 |
| Appearance | White to off-white solid powder |
| LogP | 3.5 |
| Hydrogen Bond Donor Count | 1 |
| Hydrogen Bond Acceptor Count | 9 |
| Rotatable Bond Count | 7 |
| Heavy Atom Count | 39 |
| Complexity | 962 |
| Defined Atom Stereocenter Count | 2 |
| SMILES | S1(C2C=CC=CC=2O[C@H](C)CN1CC1C(C)=CC=C(C=1)[C@@H](CC(=O)O)C1C=C(C2=C(C=1)N=NN2C)OC)(=O)=O |
| InChi Key | ZDNGJXBUEQNFBQ-XMSQKQJNSA-N |
| InChi Code | InChI=1S/C28H30N4O6S/c1-17-9-10-19(22(14-27(33)34)20-12-23-28(25(13-20)37-4)31(3)30-29-23)11-21(17)16-32-15-18(2)38-24-7-5-6-8-26(24)39(32,35)36/h5-13,18,22H,14-16H2,1-4H3,(H,33,34)/t18-,22-/m1/s1 |
| Chemical Name | (3R)-3-(7-methoxy-1-methylbenzotriazol-5-yl)-3-[4-methyl-3-[[(4R)-4-methyl-1,1-dioxo-3,4-dihydro-5,1λ6,2-benzoxathiazepin-2-yl]methyl]phenyl]propanoic acid |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| ln Vitro |
KI-696 disrupts the KEAP1-NRF2 interaction, leading to NRF2 stabilization, nuclear translocation, and activation of the antioxidant response pathway. In Normal Human Bronchial Epithelial (NHBE) cells, treatment with KI-696 for 6 hours increased nuclear NRF2 protein accumulation in a concentration-dependent manner [1]. In NHBE cells transfected with NRF2 siRNA, KI-696 (1 µM) upregulated mRNA expression of NRF2-dependent genes NQO1 and GCLM. This upregulation was significantly decreased by NRF2 gene silencing, confirming an NRF2-dependent mechanism [1]. KI-696 (1 µM) induced NQO1 specific enzyme activity by 6-fold over DMSO control in NHBE cells. NRF2 siRNA abolished approximately 88% of this induced activity [1]. In the human lung epithelial cell line BEAS-2B, KI-696 increased NQO1 enzyme activity with an EC50 of 12 nM [1]. In NHBE cells, pre-treatment with KI-696 prevented tert-butyl hydroperoxide (tBHP)-induced depletion of reduced glutathione (GSH) in a concentration-dependent manner and maintained normal cell morphology [1]. In air-liquid interface (ALI) cultures of bronchial epithelial cells derived from COPD patients, KI-696 treatment for 6 hours upregulated mRNA expression of NRF2-dependent genes (NQO1, TXNRD1, GCLM, HO-1) in a concentration-dependent manner, with an EC50 of 22 nM for NQO1 mRNA expression [1]. A fluorescence polarization (FP) assay measuring inhibition of the Kelch domain-NRF2 interaction showed that intermediate compounds during optimization (e.g., Compound 6) had an IC50 of 0.27 µM [1]. Selectivity screening against a panel of 49 in vitro functional assays revealed minor cross-reactivity with OATP1B1 (IC50 = 2.5 µM), BSEP (IC50 = 4.0 µM), and PDE3A (IC50 = 10 µM). No significant cytotoxicity was observed in BEAS-2B cells at concentrations up to 10 µM [1]. |
| ln Vivo |
In Han Wistar rats, intravenous (IV) infusion of KI-696 over 6 hours induced dose-dependent increases in the pulmonary expression of NRF2-regulated genes (Nqo1, Ho-1, Txnrd1, Srxn1, Gsta3, Gclc). The average EC50 for gene induction across these six genes was 36.4 ± 3.4 µmol/kg [1]. IV infusion of KI-696 (35 µmol/kg, approximate average EC50 for gene expression) attenuated ozone-induced pulmonary inflammation in rats. It significantly reduced the ozone-induced increases in total cells, neutrophils, and mononuclear cells in bronchoalveolar lavage (BAL) fluid [1]. IV infusion of KI-696 (10, 35, 50 µmol/kg) restored ozone-induced depletion of lung glutathione (GSH) levels in a dose-dependent manner [1]. |
| Enzyme Assay |
The inhibition of the KEAP1 Kelch domain-NRF2 interaction was determined using a fluorescence polarization (FP)-based competition assay. The assay was performed in a 384-well microplate. Each well contained a fluorescently labeled NRF2-derived peptide (5’-TAMRA-AFFAQLQLDEETGEFL) at 2 nM and the human KEAP1 Kelch domain (residues 321-609) at 7 nM in assay buffer (50 mM Tris-HCl pH 8.0, 100 mM NaCl, 5 mM MgCl2, 2 mM CHAPS, 1 mM DTT, 0.005% BSA, 1% DMSO). After incubation for 1 hour at room temperature, fluorescence polarization (excitation 485 nm/emission 520 nm) was measured. IC50 values were determined by fitting the data to a four-parameter logistic model [1]. Binding affinity (Kd) was measured by Isothermal Titration Calorimetry (ITC). Experiments were performed at 25°C in a buffer containing 50 mM HEPES-NaOH pH 7.5, 100 mM NaCl, 1 mM TCEP, and 5% DMSO. The compound was titrated into the sample cell containing the human KEAP1 Kelch domain (residues 321-609). Data were analyzed using appropriate software, and heats of dilution from the final injections were subtracted before analysis [1]. |
| Cell Assay |
NRF2 Nuclear Translocation: NHBE cells were plated and cultured until confluent. Cells were treated with compounds or DMSO for 6 hours. Cytoplasmic and nuclear fractions were generated using a modified commercial kit protocol. NRF2 protein levels in the nuclear fraction were quantified using a commercial ELISA-based kit. Data were reported as optical density per µg total protein relative to DMSO control [1]. Gene Expression (siRNA): NHBE cells were transfected with non-targeting or NRF2 siRNA for 48 hours. On the second day post-transfection, cells were treated with compounds for approximately 24 hours. Total RNA was isolated, reverse transcribed to cDNA, and quantified by real-time PCR using specific primer/probe sets for target genes (e.g., NQO1, GCLM) and a housekeeping gene (18S rRNA). Fold changes were calculated using the comparative Ct method [1]. NQO1 Enzyme Activity: NHBE cells were transfected with siRNA for 48 hours, then treated with compounds for an additional 48 hours. Cells were lysed, and protein concentration was determined. NQO1 activity in crude lysates was measured kinetically by monitoring the reduction of a substrate coupled to a colorimetric change, with and without a specific NQO1 inhibitor (dicoumarol). Specific activity was calculated by subtracting the inhibitor-treated values, normalizing to total protein, and expressed as µmol/min/mg protein [1]. BEAS-2B NQO1 Activity: BEAS-2B cells were plated in 384-well plates overnight. Compounds were added for 48 hours. Medium was aspirated, and cells were lysed. NQO1 activity in lysates was measured kinetically using an MTT-based assay, monitoring absorbance at 570 nm over time. Product formation rate was plotted against compound concentration to determine EC50 [1]. Glutathione (GSH) Assay: NHBE cells were treated with compounds for 18 hours, followed by exposure to an oxidative stressor (tBHP) or controls for 4 hours. Cells were then assayed for total glutathione content using a commercial fluorescent detection kit according to the manufacturer's protocol. Results were quantified based on a standard curve [1]. COPD Patient Cell Gene Expression: Bronchial epithelial cells from COPD patients were expanded and differentiated under air-liquid interface (ALI) culture conditions for approximately 28 days. Differentiated ALI cultures were treated with compounds for 6 hours. Total RNA was isolated and subjected to real-time PCR analysis for NRF2 target genes using specific primer/probe sets. Fold changes were calculated relative to a housekeeping gene [1]. |
| Animal Protocol |
Pharmacokinetics Study: KI-696 was administered to Han Wistar rats in a standard intravenous (IV) and oral (PO) crossover design. Blood samples were collected over 24 hours, and compound concentrations were determined by LC/MS/MS. Pharmacokinetic parameters were calculated from the blood concentration-time data [1]. In Vivo Gene Expression Study: Male Han Wistar rats received a 6-hour IV infusion of KI-696 or vehicle at various doses (targeting total delivered doses of 5, 10, 25, 50 µmol/kg). Immediately after the infusion, lungs were collected, snap-frozen, and later analyzed for mRNA expression of NRF2-regulated genes by real-time PCR [1]. Ozone-Induced Inflammation Model: Han Wistar rats were administered KI-696 or vehicle by IV infusion over 6 hours. Twenty-four hours after the start of infusion, rats were exposed to ozone (1 ppm) or air for 3 hours. Fifteen minutes post-exposure, rats were euthanized. Bronchoalveolar lavage (BAL) was performed to collect inflammatory cells for total and differential cell counts. In a separate study, lungs were collected post-ozone exposure for GSH measurement using a commercial assay kit. Blood samples were also collected during the infusion to determine compound concentrations [1]. |
| ADME/Pharmacokinetics |
Following IV administration in Han Wistar rats, KI-696 exhibited relatively high clearance and a short half-life [1]. Oral bioavailability (F) of KI-696 in rats was low, reported as 7% [1]. During a 6-hour IV infusion in rats targeting doses of 10, 35, and 50 µmol/kg, steady-state blood concentrations of KI-696 were 407 ± 44 nM, 946 ± 50 nM, and 1437 ± 186 nM, respectively [1]. |
| Toxicity/Toxicokinetics |
In an enhanced cross-screen panel (eXP) consisting of 49 in vitro functional assays for potential toxicity liabilities, KI-696 showed inhibition of OATP1B1 (IC50 = 2.5 µM), BSEP (IC50 = 4.0 µM), and PDE3A (IC50 = 10 µM). No significant cross-reactivity was observed in the other assays [1]. No cytotoxicity was observed towards BEAS-2B cells with KI-696 at concentrations up to 10 µM in a cell viability assay [1]. |
| References |
[1]. Monoacidic Inhibitors of the Kelch-like ECH-Associated Protein 1: Nuclear Factor Erythroid 2-Related Factor 2 (KEAP1:NRF2) Protein-Protein Interaction with High Cell Potency Identified by Fragment-Based Discovery. J Med Chem. 2016 Apr 28. |
| Additional Infomation |
KI-696 was discovered using a fragment-based drug discovery (FBDD) approach. X-ray crystallographic screening identified three binding "hot-spots" within the NRF2 pocket of KEAP1. Optimization from a weak fragment hit (Kd >1 mM) led to KI-696 with nanomolar affinity [1]. The compound is a mono-acidic inhibitor containing a single carboxylic acid, contrasting with multi-acidic peptide ligands. It is not a peptidomimetic but fulfills a three-point pharmacophore (acid, aromatic-acceptor, sulfonamide) identified from the fragment screen [1]. The primary proposed therapeutic application is for diseases involving oxidative stress and inflammation, such as Chronic Obstructive Pulmonary Disease (COPD). The study demonstrates its activity in cellular and animal models relevant to COPD [1]. KI-696 represents a non-covalent, direct inhibitor of the KEAP1-NRF2 PPI, which may offer advantages in selectivity over electrophilic NRF2 activators that covalently modify KEAP1 cysteines [1]. The compound is positioned as a high-quality chemical probe for exploring the therapeutic potential of disrupting the KEAP1-NRF2 interaction [1]. |
Solubility Data
| Solubility (In Vitro) | DMSO : ~125 mg/mL (~227.01 mM) |
| Solubility (In Vivo) |
Solubility in Formulation 1: ≥ 2.08 mg/mL (3.78 mM) (saturation unknown) in 10% DMSO + 40% PEG300 +5% Tween-80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 20.8 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 + to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.8161 mL | 9.0805 mL | 18.1610 mL | |
| 5 mM | 0.3632 mL | 1.8161 mL | 3.6322 mL | |
| 10 mM | 0.1816 mL | 0.9081 mL | 1.8161 mL |