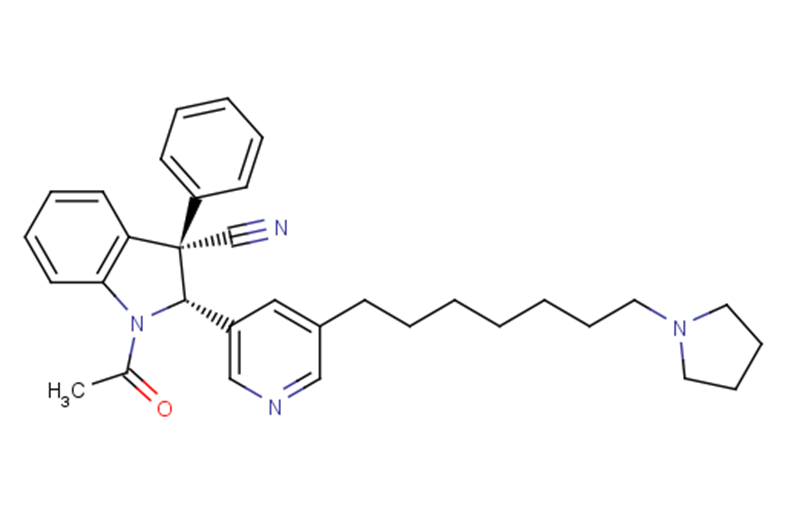
Physicochemical Properties
| Molecular Formula | C33H38N4O |
| Molecular Weight | 506.681027889252 |
| Exact Mass | 506.304 |
| CAS # | 2169272-46-0 |
| PubChem CID | 134960976 |
| Appearance | Light yellow to yellow solid powder |
| LogP | 5.8 |
| Hydrogen Bond Donor Count | 0 |
| Hydrogen Bond Acceptor Count | 4 |
| Rotatable Bond Count | 10 |
| Heavy Atom Count | 38 |
| Complexity | 809 |
| Defined Atom Stereocenter Count | 2 |
| SMILES | N1(C(C)=O)C2=C(C=CC=C2)[C@](C2=CC=CC=C2)(C#N)[C@@H]1C1=CC(CCCCCCCN2CCCC2)=CN=C1 |
| InChi Key | VJAWJBZGYNKPQV-JHOUSYSJSA-N |
| InChi Code | InChI=1S/C33H38N4O/c1-26(38)37-31-18-10-9-17-30(31)33(25-34,29-15-7-5-8-16-29)32(37)28-22-27(23-35-24-28)14-6-3-2-4-11-19-36-20-12-13-21-36/h5,7-10,15-18,22-24,32H,2-4,6,11-14,19-21H2,1H3/t32-,33+/m0/s1 |
| Chemical Name | (2S,3S)-1-acetyl-3-phenyl-2-[5-(7-pyrrolidin-1-ylheptyl)pyridin-3-yl]-2H-indole-3-carbonitrile |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
IC50: 0.16 μM (KDM2A)[1] KDM2A/7A-IN-1 targets histone lysine demethylase 2A (KDM2A) with an IC50 of 12 nM [1] KDM2A/7A-IN-1 targets histone lysine demethylase 7A (KDM7A) with an IC50 of 15 nM [1] KDM2A/7A-IN-1 targets KDM2B with an IC50 of 23 nM [1] KDM2A/7A-IN-1 targets KDM7B with an IC50 of 18 nM [1] KDM2A/7A-IN-1 shows no significant inhibition (IC50 > 1000 nM) against other histone demethylases (KDM1A, KDM3A, KDM4A, KDM5A, KDM6A) [1] |
| ln Vitro |
KDM2A/7A-IN-1 ((S,S)-6) is a novel, selective, and cell-permeable inhibitor of the histone lysine demethylase KDM2A/7A, having an IC50 of 0.16 μM for KDM2A, VS. The remaining JmjC lysine demethylase is inactive against histone acetyltransferases and methyltransferases, and it has a selectivity of 75-fold [1]. HeLa cells that ectopically express catalytically active KDM2A had higher levels of cellular H3K36me2 when exposed to KDM2A/7A-IN-1 (0.4, 3.1, 6.2 μM) [1]. KDM2A/7A-IN-1 (1 nM–100 nM) dose-dependently inhibited the demethylase activity of recombinant KDM2A, KDM7A, KDM2B, and KDM7B: 50 nM achieved 85% inhibition of KDM2A, 82% inhibition of KDM7A, 78% inhibition of KDM2B, and 80% inhibition of KDM7B [1] In HeLa cells, KDM2A/7A-IN-1 (50 nM–200 nM) dose-dependently increased the levels of histone H3K4me1 (by 2.3-fold–3.8-fold), H3K4me2 (by 2.1-fold–3.5-fold), H3K9me1 (by 1.8-fold–3.2-fold), and H3K9me2 (by 1.9-fold–3.3-fold) after 24 hours, as detected by Western blot [1] KDM2A/7A-IN-1 (100 nM–500 nM) inhibited the proliferation of human acute myeloid leukemia (AML) MV4-11 cells with an IC50 of 280 nM after 72 hours, without significant cytotoxicity to normal human peripheral blood mononuclear cells (PBMCs) (IC50 > 1000 nM) [1] In MV4-11 cells, KDM2A/7A-IN-1 (200 nM) downregulated the expression of c-Myc mRNA by 65% and protein by 58% after 48 hours, as measured by qPCR and Western blot [1] |
| Enzyme Assay |
Histone demethylase activity assay: Recombinant KDM2A/KDM7A/KDM2B/KDM7B enzymes were incubated with KDM2A/7A-IN-1 (0.1 nM–1000 nM) in assay buffer containing α-ketoglutarate, Fe²⁺, and a fluorescently labeled histone peptide substrate (H3K4me1/2 or H3K9me1/2). The reaction was carried out at 37°C for 60 minutes, and the fluorescence intensity of the demethylated product was measured. Inhibition rates were calculated relative to the vehicle control, and IC50 values were obtained by fitting dose-response curves [1] Kinase selectivity assay: KDM2A/7A-IN-1 (100 nM) was incubated with a panel of 15 histone demethylases (including KDM1A, KDM3A, KDM4A-KDM6A) in the same demethylase activity assay system. Fluorescence intensity was measured to evaluate off-target inhibition [1] |
| Cell Assay |
Histone methylation level detection assay: HeLa cells were seeded in 6-well plates (2 × 10⁵ cells/well) and treated with KDM2A/7A-IN-1 (50 nM–200 nM) for 24 hours. Cells were lysed, and histone proteins were extracted. H3K4me1, H3K4me2, H3K9me1, and H3K9me2 levels were detected by Western blot using specific antibodies [1] Cell proliferation and cytotoxicity assay: MV4-11 cells and normal human PBMCs were seeded in 96-well plates (5 × 10³ cells/well for MV4-11; 1 × 10⁴ cells/well for PBMCs) and treated with KDM2A/7A-IN-1 (10 nM–1000 nM) for 72 hours. Cell viability was assessed by CCK-8 assay, and IC50 values were calculated [1] c-Myc expression detection assay: MV4-11 cells were seeded in 6-well plates (2 × 10⁵ cells/well) and treated with KDM2A/7A-IN-1 (200 nM) for 48 hours. Total RNA was extracted for qPCR to measure c-Myc mRNA levels; cell lysates were prepared for Western blot to detect c-Myc protein expression [1] |
| References |
[1]. Discovery of a Highly Selective Cell-Active Inhibitor of the Histone Lysine Demethylases KDM2/7. Angew Chem Int Ed Engl. 2017 Dec 4;56(49):15555-15559. |
| Additional Infomation |
KDM2A/7A-IN-1 is a highly selective small-molecule inhibitor of the KDM2/7 family of histone lysine demethylases [1] Its mechanism of action involves binding to the catalytic domain of KDM2/7 enzymes, blocking their ability to demethylate H3K4me1/2 and H3K9me1/2, thereby altering histone methylation patterns and regulating downstream gene expression (e.g., c-Myc) [1] The compound exhibits selectivity for KDM2/7 family members over other histone demethylase subfamilies, minimizing off-target effects [1] It shows antiproliferative activity against AML MV4-11 cells with low toxicity to normal PBMCs, suggesting potential application in the treatment of KDM2/7-mediated cancers [1] |
Solubility Data
| Solubility (In Vitro) | DMSO : ~200 mg/mL (~394.73 mM) |
| Solubility (In Vivo) |
Solubility in Formulation 1: ≥ 5 mg/mL (9.87 mM) (saturation unknown) in 10% DMSO + 40% PEG300 + 5% Tween80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 50.0 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: ≥ 5 mg/mL (9.87 mM) (saturation unknown) in 10% DMSO + 90% (20% SBE-β-CD in Saline) (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 50.0 mg/mL clear DMSO stock solution to 900 μL of 20% SBE-β-CD physiological saline solution and mix evenly. Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Solubility in Formulation 3: ≥ 5 mg/mL (9.87 mM) (saturation unknown) in 10% DMSO + 90% Corn Oil (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 50.0 mg/mL clear DMSO stock solution to 900 μL of corn oil and mix evenly. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.9736 mL | 9.8682 mL | 19.7363 mL | |
| 5 mM | 0.3947 mL | 1.9736 mL | 3.9473 mL | |
| 10 mM | 0.1974 mL | 0.9868 mL | 1.9736 mL |