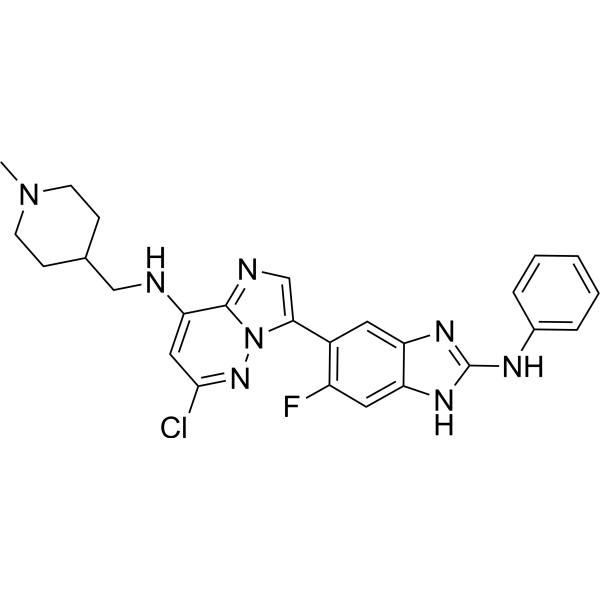
Physicochemical Properties
| Molecular Formula | C26H26CLFN8 |
| Molecular Weight | 504.99 |
| Exact Mass | 504.195 |
| CAS # | 2328097-41-0 |
| PubChem CID | 139593294 |
| Appearance | White to off-white solid powder |
| LogP | 5.3 |
| Hydrogen Bond Donor Count | 3 |
| Hydrogen Bond Acceptor Count | 7 |
| Rotatable Bond Count | 6 |
| Heavy Atom Count | 36 |
| Complexity | 720 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | FC1C(=CC2=C(C=1)NC(NC1C=CC=CC=1)=N2)C1=CN=C2C(NCC3CCN(C)CC3)=CC(=NN12)Cl |
| InChi Key | BPRAHARGXSNSSO-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C26H26ClFN8/c1-35-9-7-16(8-10-35)14-29-22-13-24(27)34-36-23(15-30-25(22)36)18-11-20-21(12-19(18)28)33-26(32-20)31-17-5-3-2-4-6-17/h2-6,11-13,15-16,29H,7-10,14H2,1H3,(H2,31,32,33) |
| Chemical Name | 3-(2-anilino-6-fluoro-1H-benzimidazol-5-yl)-6-chloro-N-[(1-methylpiperidin-4-yl)methyl]imidazo[1,2-b]pyridazin-8-amine |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| ln Vitro | In human cells, IRE1α kinase-IN-1 (Compound 31) suppresses endoribonuclease activity and stops IRE1α oligomerization and phosphorylation caused by endoplasmic reticulum stress [1]. With only 4 out of 455 kinases showing >70% inhibition, IRE1α kinase-IN-1 is incredibly selective. With an IC50 of 160 nM, IRE1α kinase-IN-1 prevents autophosphorylation of the recombinant G547 IRE1α KEN domain pS274. With an IC50 of 0.74 μM, IRE1α kinase-IN-1 suppresses tunicamycin-induced GFP-IRE1α foci in HEK293 cells. With an IC50 of 0.27 μM, IRE1α kinase-IN-1 prevents ATP site LanthaScreen tracer from binding to recombinant dephosphorylated G547 IRE1α KEN [1]. With an IC50 range of 0.68-1.63 μM, IRE1α kinase-IN-1 suppresses XBP1 luciferase fusion mRNA splicing in HEK293 cells that is stimulated by thapsigargin and tunicamycin [1]. In NCI-H929 cells, IRE1α kinase-IN-1 (0 – 20 μM) dose-dependently suppresses tunicamycin-induced XBP1 expression. IRE1α kinase-IN-1 (0 – 20 μM) reduces IRE1α-dependent XBP1s mRNA expression in H929 cells. inside cells [1]. |
| References |
[1]. Binding to an Unusual Inactive Kinase Conformation by Highly Selective Inhibitors of Inositol-Requiring Enzyme 1α Kinase-Endoribonuclease. J Med Chem. 2019;62(5):2447-2465. |
Solubility Data
| Solubility (In Vitro) | DMSO: 25 mg/mL (49.51 mM) |
| Solubility (In Vivo) |
Solubility in Formulation 1: ≥ 2.5 mg/mL (4.95 mM) (saturation unknown) in 10% DMSO + 40% PEG300 + 5% Tween80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 25.0 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: ≥ 2.5 mg/mL (4.95 mM) (saturation unknown) in 10% DMSO + 90% Corn Oil (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 25.0 mg/mL clear DMSO stock solution to 900 μL of corn oil and mix evenly. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.9802 mL | 9.9012 mL | 19.8024 mL | |
| 5 mM | 0.3960 mL | 1.9802 mL | 3.9605 mL | |
| 10 mM | 0.1980 mL | 0.9901 mL | 1.9802 mL |