H3B-8800 is a novel, potent and orally bioavailable modulator of the SF3b complex. H3B-8800 potently and preferentially kills spliceosome-mutant epithelial and hematologic tumor cells. These killing effects of H3B-8800 are due to its direct interaction with the SF3b complex, as evidenced by loss of H3B-8800 activity in drug-resistant cells bearing mutations in genes encoding SF3b components. Although H3B-8800 modulates WT and mutant spliceosome activity, the preferential killing of spliceosome-mutant cells is due to retention of short, GC-rich introns, which are enriched for genes encoding spliceosome components. These data demonstrate the therapeutic potential of splicing modulation in spliceosome-mutant cancers. Genomic analyses of cancer have identified recurrent point mutations in the RNA splicing factor-encoding genes SF3B1, U2AF1, and SRSF2 that confer an alteration of function. Cancer cells bearing these mutations are preferentially dependent on wild-type (WT) spliceosome function, but clinically relevant means to therapeutically target the spliceosome do not currently exist.
Physicochemical Properties
| Molecular Formula | C31H45N3O6 |
| Molecular Weight | 555.705508947372 |
| Exact Mass | 555.33 |
| Elemental Analysis | C, 67.00; H, 8.16; N, 7.56; O, 17.27 |
| CAS # | 1825302-42-8 |
| Related CAS # | 1825302-42-8; |
| PubChem CID | 92135969 |
| Appearance | Yellow to orange solid |
| Density | 1.2±0.1 g/cm3 |
| Boiling Point | 710.6±60.0 °C at 760 mmHg |
| Flash Point | 383.6±32.9 °C |
| Vapour Pressure | 0.0±2.4 mmHg at 25°C |
| Index of Refraction | 1.581 |
| LogP | 2.22 |
| Hydrogen Bond Donor Count | 2 |
| Hydrogen Bond Acceptor Count | 8 |
| Rotatable Bond Count | 6 |
| Heavy Atom Count | 40 |
| Complexity | 928 |
| Defined Atom Stereocenter Count | 6 |
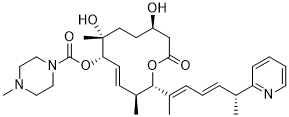
| SMILES | O(C(N1CCN(C)CC1)=O)[C@H]1C=C[C@H](C)[C@@H](/C(=C/C=C/[C@@H](C)C2C=CC=CN=2)/C)OC(C[C@@H](CC[C@@]1(C)O)O)=O |t:12| |
| InChi Key | YOIQWBAHJZGRFW-WVRLKXNASA-N |
| InChi Code | InChI=1S/C31H45N3O6/c1-22(26-11-6-7-16-32-26)9-8-10-23(2)29-24(3)12-13-27(39-30(37)34-19-17-33(5)18-20-34)31(4,38)15-14-25(35)21-28(36)40-29/h6-13,16,22,24-25,27,29,35,38H,14-15,17-21H2,1-5H3/b9-8+,13-12+,23-10+/t22-,24+,25-,27+,29-,31-/m1/s1 |
| Chemical Name | (2S,3S,6S,7R,10R,E)-7,10-Dihydroxy-3,7-dimethyl-12-oxo-2-((R,2E,4E)-6-(pyridin-2-yl)hepta-2,4-dien-2-yl)oxacyclododec-4-en-6-yl 4-methylpiperazine-1-carboxylate |
| Synonyms | H3B-8800; H3B8800; RVT-2001; UNII-90YLS47BRX; RVT2001; 1825302-42-8; 90YLS47BRX; 1-Piperazinecarboxylic acid, 4-methyl-, (2S,3S,4E,6S,7R,10R)-7,10-dihydroxy-3,7-dimethyl-2-((1E,3E,5R)-1-methyl-5-(2-pyridinyl)-1,3-hexadien-1-yl)-12-oxooxacyclododec-4-en-6-yl ester; SCHEMBL17255784; EX-A8015; H3B 8800 [WHO-DD]; H3B 8800 |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
SF3B splicing SF3b complex (spliceosome) [1] The compound competes with pladienolide for binding to SF3b complexes containing either wild-type or mutant SF3B1. [1] |
| ln Vitro |
H3B-8800 potently and preferentially kills spliceosome-mutant epithelial and hematologic tumor cells. These killing effects of H3B-8800 are due to its direct interaction with the SF3b complex, as evidenced by loss of H3B-8800 activity in drug-resistant cells bearing mutations in genes encoding SF3b components. H3B-8800 potently binds to and inhibits the splicing activity of both wild-type and mutant SF3B1-containing spliceosomes in competitive binding assays. [1] It induces dose-dependent inhibition of both canonical and aberrant splicing in biochemical splicing assays using nuclear extracts. [1] In a panel of pancreatic cancer cell lines, H3B-8800 treatment for 72 hours resulted in cellular lethality and caspase-3/7 cleavage specifically in the SF3B1-mutant cell line Panc05.04, but not in wild-type lines. [1] It demonstrated preferential killing of K562 cells bearing the SF3B1-K700E mutation compared to isogenic wild-type counterparts. [1] Treatment with H3B-8800 modulated both canonical and aberrant splicing in Panc05.04 cells, as shown by decreased mature MBD4 mRNA levels, accumulation of MBD4 pre-mRNA, and decreased mutant SF3B1-induced aberrant splicing of MAP3K7. [1] It inhibited ATP-dependent 17S U2 snRNP complex formation in nuclear extracts, indicating it interferes with precatalytic spliceosome assembly by disrupting the interaction between the SF3b complex and the branchpoint region. [1] Resistance mutations in SF3B1 (R1074H) or PHF5A (Y36C) conferred resistance to H3B-8800, confirming its selectivity for the SF3b complex. [1] RNA-seq analysis showed that H3B-8800 treatment preferentially induced retention of short, GC-rich introns, particularly in genes encoding spliceosome components such as U2AF2 and RBM10. [1] It induced a dose-dependent reduction in U2AF2 and SRPK1 protein expression in K562 cells. [1] |
| ln Vivo |
Oral administration of 2 or 4 mg H3B-8800 per kg body weight (mg/kg) daily slowed the growth of xenografts with SF3B1K700E (P < 0.003, two-way ANOVA followed by Dunnett’s multiple comparison test) but had no effect on SF3B1WT xenografts. Although a dose of 8 mg/kg did slow the growth of SF3B1WT tumors, it completely abrogated growth of SF3B1K700E tumors (P < 0.003).b). H3B-8800 similarly demonstrated significant antitumor activity against xenografts of human HNT-34 acute myeloid leukemia (AML) cells bearing the endogenous mutation encoding SF3B1-K700E. H3B-8800 reached similar concentrations in plasma as well as in tumors of mice harboring SF3B1WT or SF3B1K700E xenografts, and H3B-8800 modulated both canonical and aberrant splicing in a dose-dependent manner. Splicing modulation was observed as early as 1 h following treatment, and splicing returned to pretreatment levels within 24 h following treatment, in accordance with the H3B-8800’s pharmacokinetic profile. In NSG mice bearing xenografts of isogenic K562 cells, oral administration of H3B-8800 (2 or 4 mg/kg daily) significantly slowed tumor growth in SF3B1-K700E mutants but had no effect on wild-type tumors. A dose of 8 mg/kg completely abrogated growth of SF3B1-K700E tumors and also slowed growth of wild-type tumors. [1] H3B-8800 demonstrated significant antitumor activity against xenografts of human HNT-34 AML cells bearing endogenous SF3B1-K700E mutation. [1] In mice bearing AML patient-derived xenografts (PDXs) with SF3B1 mutation, 10-day treatment with H3B-8800 (8 mg/kg) significantly reduced leukemic burden, while having negligible effects in wild-type PDX recipients. [1] Similar preferential activity was observed in CMML PDX models with SRSF2 mutations, where H3B-8800 treatment reduced human leukemic burden in peripheral blood and organs, and reduced spleen and liver size only in SRSF2-mutant models. [1] Splicing modulation (reduction of mature mRNA and accumulation of pre-mRNA) was observed in tumors as early as 1 hour post-treatment and returned to baseline within 24 hours. [1] |
| Enzyme Assay |
SF3B complexes were immunoprecipitated from nuclear extracts prepared from 293F cells overexpressing Flag-tagged SF3B1. First, batch immobilization of antibody to beads was performed through incubating 80 μg of anti-SF3B1 antibody (MBL International D221–3, Anti-Sap155 monoclonal antibody) and 24 mg anti-mouse PVT scintillation proximity assay (SPA) beads (PerkinElmer) for 30 min. After centrifugation (14,000 r.p.m. for 5 min at 4 °C), the antibody–bead mixture was resuspended in PBS supplemented with PhosSTOP phosphatase inhibitor cocktail (Roche) and cOmplete ULTRA protease inhibitor cocktail (Roche). Nuclear extracts were prepared through diluting 40 mg into a total volume of 16 ml PBS with phosphatase and protease inhibitors, and the mixture was centrifuged (14,000 r.p.m. for 10 min at 4 °C). The supernatant was transferred into a clean tube, and the antibody–bead mixture was added and incubated for 2 h. The beads were collected via centrifuging, washed twice with PBS + 0.1% Triton X-100, and resuspended with 4.8 ml of PBS. 100 μl binding reactions were prepared using slurry and varying concentrations of H3B-8800. After 15 min preincubation at room temperature, 1 nM 3H-probe used in Kotake et al.20 was added. The mixture was incubated at room temperature for 15 min, and luminescence signals were read using a MicroBeta2 Plate Counter (PerkinElmer). Competitive binding assays were performed using SF3b complexes immunoprecipitated from nuclear extracts of cells overexpressing Flag-tagged SF3B1. Anti-SF3B1 antibody was immobilized on beads and incubated with nuclear extracts. Binding reactions were set up with bead slurry and varying concentrations of H3B-8800, followed by addition of a radiolabeled probe. Luminescence signals were measured to determine competition with pladienolide binding. [1] An in vitro splicing reaction assay was performed using nuclear extracts and pre-mRNA substrates (Ad2-derived or Ad2-ZDHHC16 hybrid). Reactions containing nuclear extract, pre-mRNA, control RNA, and varying concentrations of H3B-8800 or DMSO were preincubated, then splicing activation buffer (containing ATP and MgCl2) was added. After incubation, reactions were quenched and analyzed by RT-qPCR to quantify canonical and aberrant splicing junctions. [1] A U2 snRNP complex assembly assay was performed using nuclear extracts depleted of ATP. Extracts were preincubated with or without H3B-8800, then incubated with ATP, MgCl2, creatine phosphate, and a radiolabeled 2′-O-methyl oligonucleotide complementary to the U2 snRNA branchpoint binding region. Samples were purified and run on native agarose gels to resolve 17S and 12S U2 snRNP complexes. [1] |
| Cell Assay |
Pancreatic cancer cells were seeded at 750 cells per well in a 384 well plate and treated with compound on day 2. K562 isogenic cells (K562-SF3B1K700E and K562-SF3B1K700K) were seeded at 10,000 cells per well in 96-well plates and treated with compound 4 h later. The relative numbers of viable and apoptotic cells were measured via luminescence using CellTiter-Glo or Caspase-Glo 3/7 (Promega) at the indicated time points as instructed. Cell viability and apoptosis assays were performed using pancreatic cancer cells or K562 isogenic cells seeded in multi-well plates. Cells were treated with H3B-8800 or vehicle, and viability/apoptosis was measured at indicated time points using luminescence-based assays (CellTiter-Glo or Caspase-Glo 3/7). [1] For splicing analysis, cells were treated with H3B-8800 for 6 hours, then RNA was extracted and reverse transcribed. Splicing events were quantified using TaqMan Low-Density Arrays (TLDA) or RT-qPCR with specific primer-probe sets for targets such as MBD4, MAP3K7, U2AF2, and aberrant junctions. [1] Western blotting was performed on cell lysates from treated K562 isogenic cells. Proteins were separated by SDS-PAGE, transferred to membranes, and probed with antibodies against U2AF2, SRPK1, MCL1, and loading controls (β-actin or GAPDH). [1] RNA-seq sample preparation involved treating NALM-6 or K562 isogenic cells with H3B-8800 for 6 hours, followed by RNA isolation, cDNA library preparation, and sequencing. Data were analyzed for differential splicing events, intron retention, and gene expression changes. [1] |
| Animal Protocol |
K562 isogenic lines, HNT-34 and K052 xenograft models efficacy and pharmacokinetic/pharmacodynamic modeling. 1 × 107 K562 isogenic, HNT-34, or K052 cells were subcutaneously implanted into the flank of female NSG or CB17-SCID mice of 6–8 weeks of age. Mice were treated with H3B-8800 (10% ethanol, 5% Tween-80, 85% saline) or vehicle control. For the efficacy studies, the mice were orally dosed daily, and the mice were monitored until they reached either of the following endpoints: (i) excessive tumor volume (≥20 mm in its longest diameter), which was measured three times a week (tumor volume calculated by using the ellipsoid formula: (length × width2) / 2), or (ii) development of any health problem, such as paralysis or excessive body weight loss. The differences in tumor volume during the study period between the vehicle-treated and H3B-8800-treated groups were analyzed by two-way analysis of variance (ANOVA) followed by the Dunnett’s multiple comparison test. For the pharmacokinetic/pharmacodynamic modeling (PK/PD) studies, the mice were treated with one dose, and the tumors were collected at the indicated times (in Figs. 2 and and3)3) after treatment for further analysis. RNA was isolated using RiboPure RNA purification kit (Ambion) and used for TLDA. All mouse studies were carried out under protocols approved by the Institutional Animal Care and Use Committee at H3 Biomedicine. For xenograft efficacy studies, female NSG or CB17-SCID mice (6-8 weeks old) were subcutaneously implanted with tumor cells (e.g., K562 isogenic, HNT-34, or K052 cells). Mice were orally dosed daily with H3B-8800 (formulated in 10% ethanol, 5% Tween-80, 85% saline) or vehicle control. Tumor volume was measured regularly, and mice were monitored until reaching endpoint criteria (excessive tumor size or health issues). [1] For pharmacokinetic/pharmacodynamic (PK/PD) studies, mice received a single oral dose of H3B-8800, and tumors were collected at specified time points for RNA analysis (e.g., TLDA) to assess splicing modulation. [1] Patient-derived xenograft (PDX) models were established by transplanting patient-derived hematopoietic cells into sublethally irradiated NSGS mice. Once engraftment was confirmed (human CD45+ cells >3% in peripheral blood or hemoglobin ≤11 g/dL), mice were randomized to receive vehicle or H3B-8800 (8 mg/kg orally daily for 10 days). Mice were euthanized 2 hours after the last dose for analysis of leukemic burden and organ infiltration. [1] |
| ADME/Pharmacokinetics |
H3B-8800 reached similar concentrations in plasma and tumors of mice bearing either SF3B1 wild-type or mutant xenografts after oral administration. [1] Splicing modulation in tumors was observed as early as 1 hour post-dose and returned to pretreatment levels within 24 hours, consistent with its pharmacokinetic profile. [1] |
| References |
[1]. H3B-8800, an orally available small-molecule splicing modulator, induces lethality in spliceosome-mutant cancers. Nat Med. 2018 May;24(4):497-504. |
| Additional Infomation |
H3B-8800 is a novel spliceosome inhibitor developed by H3 Biomedicine. It offers the benefit of preferentially killing spliceosome-mutant cancer cells whereas other splicesome inhibitors, such as the pladienolide analogue E7107, show no such preferential targeting. H3B-8800 was granted orphan drug status by the FDA in August 2017 and is in clinical trials for the treatment of acute myelogenous leukemia and chronic myelomonocytic leukemia. Splicing Inhibitor H3B-8800 is an orally bioavailable inhibitor of the splicing factor 3B subunit 1 (SF3B1), with potential antineoplastic activity. Upon administration, H3B-8800 binds to and blocks the activity of SF3B1, a core spliceosome protein that is mutated in various cancer cells. This modulates RNA splicing by preventing aberrant mRNA splicing by the spliceosome, blocks RNA mis-splicing, enhances proper RNA splicing and prevents the expression of certain tumor-associated genes. This leads to an induction of apoptosis and prevents tumor cell proliferation. In many cancer cells, core spliceosome proteins, including SF3B1, U2 small nuclear ribonucleoprotein auxiliary factor 1 (U2AF1), serine/arginine-rich splicing factor 2 (SRSF2) and U2 small nuclear ribonucleoprotein auxiliary factor subunit-related protein 2 (ZRSR2), are mutated and aberrantly activated leading to a dysregulation of mRNA splicing. Mechanism of Action H3B-8800 is thought to bind to a site similar to pladienolides on the SF3B complex within the spliceosome. Once bound it induces increased retention of short (<300 nucleotide) GC-rich introns through modulation of pre-mRNA processing. These intron-retained mRNA sequences are then thought to be destroyed through the nonsense-mediated decay pathway. It has been suggested that modulation by H3B-8800 is mediated by disruption of branchpoint sequence recognition by the SF3B complex as there is overall less preference for adenosine as the branchpoint nucleotide and a greater amount of sequences with weaker association to the SFB3 in introns retained with H3B-8800. It was found that 41 of 404 genes encoding spliceosome proteins contained GC-rich sequences whose retention was induced by H3B-8800. It is suggested that this is key to the specificity of H3B-8800's lethality as cells with spliceosome-mutant cells are dependent on the expression of wild-type spliceosome components for survival. Since cancer cells, as in myelodysplasia, experience SF3B1 mutations much more frequently than host cells, this allows H3B-8800 to be used to preferentially target these cells by inducing intron-retention in critical spliceosome component pre-mRNA leading to destruction of the now nonsense mature RNA ultimately cell-death due to the lack of these critical proteins. Pharmacodynamics H3B-8800 preferentially targets cells with spliceosome complexes containing mutant splicing factor 3B1 (SF3B1) protein, modulating intron splicing leading to increased death in cancer cells while having little effect on the viability cells with wild-type SF3B1. Both normal and aberrant mature mRNA are suppressed in mutant and wild-type cells, the selectivity of the lethal effect is thought to be due to the presence of mutant SF3B1 and its implications rather than a change in mechanism or potency of effect on the mutant protein over the wild-type [A32749 . Since SF3B1 is frequently mutated in cancer, this allows preferential targeting of cancer cells over host cells. H3B-8800 is an orally available small-molecule splicing modulator that selectively targets the SF3b complex of the spliceosome. [1] It induces preferential lethality in cancer cells bearing mutations in spliceosome genes (e.g., SF3B1, SRSF2), which are commonly found in myelodysplastic syndromes, leukemias, and some solid tumors. [1] Its mechanism involves preferential retention of short, GC-rich introns, particularly in genes encoding spliceosome components, thereby exacerbating the splicing deficiency already present in mutant cells and leading to selective cell death. [1] It exhibits distinct sequence-specific splicing modulation properties compared to other splicing modulators like E7107. [1] The study supports the clinical potential of H3B-8800 for treating genetically defined cancers with RNA splicing factor mutations. [1] |
Solubility Data
| Solubility (In Vitro) | DMSO: ≥ 100 mg/mL (~180 mM) |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.7995 mL | 8.9975 mL | 17.9950 mL | |
| 5 mM | 0.3599 mL | 1.7995 mL | 3.5990 mL | |
| 10 mM | 0.1800 mL | 0.8997 mL | 1.7995 mL |