GNE-6640, an analog of GNE-6776, is a novel potent and selective inhibitor of USP7 (ubiquitin-specific protease-7). It induces tumour cell death and enhances cytotoxicity with chemotherapeutic agents and targeted compounds, including PIM kinase inhibitors. Structural studies reveal that GNE-6640 non-covalently targets USP7 12 Å distant from the catalytic cysteine. GNE-6640 attenuates ubiquitin binding and thus inhibits USP7 deubiquitinase activity. GNE-6640 interacts with acidic residues that mediate hydrogen-bond interactions with the ubiquitin Lys48 side chain, suggesting that USP7 preferentially interacts with and cleaves ubiquitin moieties that have free Lys48 side chains. Deubiquitinase enzymes cleave ubiquitin from substrates and are implicated in disease; for example, ubiquitin-specific protease-7 (USP7) regulates stability of the p53 tumour suppressor and other proteins critical for tumour cell survival.
Physicochemical Properties
| Molecular Formula | C20H18N4O | |
| Molecular Weight | 330.383123874664 | |
| Exact Mass | 330.15 | |
| Elemental Analysis | C, 72.71; H, 5.49; N, 16.96; O, 4.84 | |
| CAS # | 2009273-67-8 | |
| Related CAS # |
|
|
| PubChem CID | 122531786 | |
| Appearance | White to off-white solid powder | |
| LogP | 3.8 | |
| Hydrogen Bond Donor Count | 3 | |
| Hydrogen Bond Acceptor Count | 4 | |
| Rotatable Bond Count | 3 | |
| Heavy Atom Count | 25 | |
| Complexity | 439 | |
| Defined Atom Stereocenter Count | 0 | |
| InChi Key | ZHYXJQQBKROZDX-UHFFFAOYSA-N | |
| InChi Code | InChI=1S/C20H18N4O/c1-2-16-17(13-5-8-18-14(9-13)10-23-24-18)11-22-20(21)19(16)12-3-6-15(25)7-4-12/h3-11,25H,2H2,1H3,(H2,21,22)(H,23,24) | |
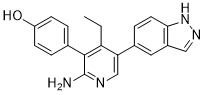
| Chemical Name | 4-[2-amino-4-ethyl-5-(1H-indazol-5-yl)pyridin-3-yl]phenol | |
| Synonyms |
|
|
| HS Tariff Code | 2934.99.03.00 | |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
|
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
USP7 (IC50 = 0.75 μM); USP7 (IC50 = 0.43 μM); USP43 (IC50 = 20.3 μM); Ub-MDM2 (IC50 = 0.23 μM) GNE-6640 encourages the natural ubiquitination of MDM2 through Lys48 (K48)-linked polyubiquitin chains, thereby guiding the process of proteasomal degradation13. GNE-6640 focuses on the p53, MDM2, and USP7 signaling pathways in cells. With an IC50 ≤ 10 μM, GNE-6640 reduces the viability of 108 cell lines. Combining GNE-6640 with DNA-damaging drugs like doxorubicin or cisplatin may trigger the p53 response and improve the effectiveness of USP7 inhibitors. GNE-6640 may cause the death of tumor cells. PIM kinase inhibitors and other targeted substances, such as chemotherapeutic agents, increase the cytotoxicity of GNE-6640[1]. |
| ln Vitro |
GNE-6640 encourages the natural ubiquitination of MDM2 through Lys48 (K48)-linked polyubiquitin chains, thereby guiding the process of proteasomal degradation13. GNE-6640 focuses on the p53, MDM2, and USP7 signaling pathways in cells. With an IC50 ≤ 10 μM, GNE-6640 reduces the viability of 108 cell lines. Combining GNE-6640 with DNA-damaging drugs like doxorubicin or cisplatin may trigger the p53 response and improve the effectiveness of USP7 inhibitors. GNE-6640 may cause the death of tumor cells. PIM kinase inhibitors and other targeted substances, such as chemotherapeutic agents, increase the cytotoxicity of GNE-6640[1]. GNE‑6640 selectively inhibits USP7 over USP47 (IC₅₀ >200 µM) and USP5 (IC₅₀ >200 µM). In cellular assays, it enhances endogenous MDM2 ubiquitination (EC₅₀ ≈ 0.23 µM in Ub‑MDM2 MSD assay) and promotes K48‑linked polyubiquitination of MDM2. It stabilizes p53 and upregulates p21 in wild‑type cells but not in p53‑null or USP7‑null cells. The compound decreases viability of 108 out of 441 cell lines (3‑day assay) with IC₅₀ ≤10 µM; in a 5‑day assay, 54 cell lines showed IC₅₀ ≤10 µM. Acute myeloid leukemia (AML) cell lines exhibited increased sensitivity to GNE‑6640. Combination with DNA‑damaging agents (e.g., doxorubicin, cisplatin) enhances its efficacy. GNE‑6640 also shows synergy with PIM kinase inhibitors (e.g., GDC‑0339, GDC‑0570) in Bliss analysis.[1] |
| Enzyme Assay |
GNE-6640, another novel selective and non-covalent inhibitor of USP7, has IC50 values of 0.75 μM (full length USP7), 0.43 μM (USP7 catalytic domain), and 20.3 μM (full length USP43). Ubiquitin-specific protease-7 (USP7) controls the stability of the p53 tumor suppressor and various other proteins critical for tumor cell survival. GNE-6640 facilitates endogenous MDM2 ubiquitination and targets cellular USP7 and p53 signaling pathways, inducing tumor cell death, enhancing cytotoxicity, and decreasing the viability of around 108 cell lines with IC50 ≤ 10 μM with chemotherapeutic agents such as PIM kinase inhibitors[2]. USP7 enzymatic analysis[1] Michaelis–Menten kinetic measurements with full-length USP7 were performed using 1 nM USP7 with a series of ubiquitin–AMC substrate titrations. The initial rate of substrate hydrolysis was determined using the Magellan software on a Tecan Safire2 plate reader, and kinetic parameters modelled using nonlinear regression analysis with GraphPad Prism software. Standard error was calculated from three technical replicates. For studies using the USP7 D305/E308 mutant, samples were reacted in a buffer consisting of 50 mM HEPES (pH 7.5), 100 mM NaCl, 2.5 mM dithiothreitol, and 0.1% (w/v) bovine gamma globulin. The starting substrate concentration of ubiquitin-Rho110 used for the Michaelis–Menten analysis was 100 μM serial diluted to 781 nM. Reactions were performed for 1 h at room temperature with a final enzyme concentration of 100 nM (three independent experiments, see symbols in plots), in black 100-μl volume 96-well half area plates. The enzymatic activity was calculated by fitting the data using the initial velocity with the linear V0 values measured by analysing the fluorescence signal of cleaved Rho-110 using excitation at 485 nm and emission at 535 nm. Deubiquitinase selectivity analysis[1] Recombinant deubiquitinase di-ubiquitin mass spectrometry cleavage assay. The MALDI–TOF DUB assay was performed using the indicated concentrations of recombinant deubiquitinases, di-ubiquitin substrates, and USP7 inhibitor compounds as described previously. The inhibition efficiency for GNE-6640 and GNE-6776 against the UCHl family members was monitored on Ub-Ube2W (Ub-E2), an alternative substrate to di-ubiquitin. USP7 activity was measured using a TR‑FRET‑based assay with GST‑UbA10‑His as substrate. The assay buffer contained HEPES (pH 7.5), detergent, DTT, and carrier protein. Compounds were pre‑incubated with recombinant full‑length USP7, then substrate was added. After reaction, detection antibodies (anti‑His‑europium and anti‑GST‑d2) were used to measure TR‑FRET signal. Inhibition was calculated relative to controls, and IC₅₀ values were determined by curve‑fitting. A di‑ubiquitin FRET assay with internally quenched K63‑linked di‑ubiquitin substrate was used as an orthogonal activity assay. Fluorescence increase upon cleavage was monitored. A ubiquitin‑rhodamine‑110 assay was also employed, where cleavage releases rhodamine‑110 and increases fluorescence. Di‑ubiquitin cleavage was further validated by mass spectrometry, monitoring conversion to ubiquitin using MRM.[1] |
| Cell Assay |
Tumour cell-line panel viability. [1] GNE-6640 and GNE-6641 were profiled for 3 days across 441 cell lines, and GNE-6776, GNE-6640, and GNE-6641 were profiled for 5 days across a subset of 185 cell lines as previously described26. In brief, compounds were screened in nine-point dose–response using a threefold dilution. Cells were seeded into 384-well plates 24 h before compound addition. Cells were then incubated with compound for 72 h or 120 h before assaying viability. Assays were performed in biological triplicate. Cells were incubated (37 °C, 5% CO2) in RPMI-1640, 2.5% FBS (72 h assay) or 5% FBS (120 h assay), and 2 mM glutamine throughout the assay. The reported IC50 and mean viability metrics were as follows: IC50 was the dose at which the estimated inhibition was 50% relative to untreated wells (that is, absolute IC50). Primary combination screen. [1] A compound library comprising 589 compounds arrayed in nine-point dose–response was screened in the absence or presence of fixed doses of GNE-6776 (0 nm, 125 nM, 250 nM, 500 nM, 1,000 nM, and 2,000 nM) or GNE-6640 (400 nM). In brief, 5,000 EOL-1 cells were seeded into 384-well plates, and compound was added 24 h later. Cell viability was determined 120 h after compound addition (CellTiter-Glo). Curves were fitted, and both IC50 and mean viability metrics were calculated. The IC50 was the dose at which inhibition was 50% relative to untreated wells. The mean viability was the average of the fitted viabilities at each tested dose. Mean viability was equivalent to the area under the log-dose/viability curve divided by the total number of tested doses. Mean viability values were used for the analysis described in Extended Data Fig. 6g. All data were fitted using Genedata Screener software. Primary combination screen analysis. [1] Normalized mean viabilities were determined in the EOL-1 cell line for 574 compounds that have known protein or mechanistic targets, in the presence of DMSO or increasing concentrations of GNE-6776 (100 nM, 250 nM, 500 nM, 1,000 nM or 2,000 nM) or 400 nM of GNE-6640. For each compound, we assessed the difference in mean viability between USP7 inhibitor treatment and the DMSO treatment. For targets targeted by three or more compounds, we calculated the enrichment of high mean viability difference for each concentration of USP7 inhibitor by using a Wilcoxon rank-sum test. For visualization purposes, we combined the results of all concentrations by taking the mean of the −log10(transformed P values) for each target. For ubiquitin‑MDM2 assays, HCT116 or SJSA‑1 cells were seeded in 96‑well plates, treated with compound for 24 h, then exposed to proteasome inhibitor MG132 for 1 h. Cells were lysed, and ubiquitinated‑MDM2 and total MDM2 were quantified using a multiplexed mesoscale immunoassay. Total MDM2 immunofluorescence: HCT‑116 cells were seeded in 384‑well plates, treated with compound for 24 h, fixed, permeabilized, stained with anti‑MDM2 primary and AlexaFluor‑555‑conjugated secondary antibodies, and imaged using a high‑content imager. Nuclear MDM2 intensity was quantified. Cell viability (IncuCyte): Cells were treated with compound in low‑serum medium containing caspase‑3/7 reagent, imaged every 2 h for 68–72 h, and confluence/caspase activity were analyzed. CellTiter‑Glo assays were performed after 72 h or 120 h of compound treatment to assess viability. Combination screens with 589 chemotherapeutic/targeted agents were conducted in EOL‑1 cells, and synergy was evaluated using Bliss analysis.[1] |
| ADME/Pharmacokinetics |
In‑vitro pharmacokinetic parameters for GNE‑6640: molecular weight 330.38 Da, logD₇.₄ 4.40, total polar surface area 87.0 Ų. Hepatic microsomal stability (CLₕₑₚ): human 14 mL/min/kg, rat 46, mouse 75, dog 24, cynomolgus 33. Hepatocyte stability (CLₕₑₚ): human 16 mL/min/kg, rat 45, mouse 68, dog 24, cynomolgus 39. Plasma protein binding: human 99.6%, rat 98.7%, mouse 99.0%. MDCK permeability: B‑to‑A 12.17 × 10⁻⁶ cm/s, A‑to‑B 14.7 × 10⁻⁶ cm/s, ratio 0.83.[1] |
| References |
[1]. USP7 small-molecule inhibitors interfere with ubiquitin binding. Nature. 2017 Oct 26;550(7677):534-538. [2]. The role of deubiquitinating enzymes in cancer drug resistance. Cancer Chemother Pharmacol. 2020 Apr;85(4):627-639 |
| Additional Infomation |
The ubiquitin system regulates essential cellular processes in eukaryotes. Ubiquitin is ligated to substrate proteins as monomers or chains and the topology of ubiquitin modifications regulates substrate interactions with specific proteins. Thus ubiquitination directs a variety of substrate fates including proteasomal degradation. Deubiquitinase enzymes cleave ubiquitin from substrates and are implicated in disease; for example, ubiquitin-specific protease-7 (USP7) regulates stability of the p53 tumour suppressor and other proteins critical for tumour cell survival. However, developing selective deubiquitinase inhibitors has been challenging and no co-crystal structures have been solved with small-molecule inhibitors. Here, using nuclear magnetic resonance-based screening and structure-based design, we describe the development of selective USP7 inhibitors GNE-6640 and GNE-6776. These compounds induce tumour cell death and enhance cytotoxicity with chemotherapeutic agents and targeted compounds, including PIM kinase inhibitors. Structural studies reveal that GNE-6640 and GNE-6776 non-covalently target USP7 12 Å distant from the catalytic cysteine. The compounds attenuate ubiquitin binding and thus inhibit USP7 deubiquitinase activity. GNE-6640 and GNE-6776 interact with acidic residues that mediate hydrogen-bond interactions with the ubiquitin Lys48 side chain, suggesting that USP7 preferentially interacts with and cleaves ubiquitin moieties that have free Lys48 side chains. We investigated this idea by engineering di-ubiquitin chains containing differential proximal and distal isotopic labels and measuring USP7 binding by nuclear magnetic resonance. This preferential binding protracted the depolymerization kinetics of Lys48-linked ubiquitin chains relative to Lys63-linked chains. In summary, engineering compounds that inhibit USP7 activity by attenuating ubiquitin binding suggests opportunities for developing other deubiquitinase inhibitors and may be a strategy more broadly applicable to inhibiting proteins that require ubiquitin binding for full functional activity.[1] \n\nHerein we describe GNE-6640 and GNE-6776, selective USP7 inhibitors that possess a structurally defined mechanism of inhibition. Establishing stringent screening cascades was critical for selecting and optimizing on-target inhibitors. Combination studies revealed a previously undescribed intersection between USP7 deubiquitinase activity and PIM kinases in regulating cell viability. Co-crystal structures of GNE-6640 or GNE-6776 pointed to the importance of the complementary charged interactions between USP7-D305/E308 and ubiquitin-K48 side chains, which we confirmed using mutational analysis. Notably, D305G has been identified as a somatic loss-of-function mutant in patients with acute lymphoblastic leukaemia21. NMR analysis of USP7 binding to native mono-ubiquitin and differentially labelled di-ubiquitins revealed that USP7 preferentially interacts with ubiquitin moieties having free K48 side chains. It has been proposed that the inefficiency of some deubiquitinases to depolymerize longer substrate-conjugated K48-linked chains enables a threshold for proteasome-targeting polyubiquitination22; our studies substantiate this idea and provide a biophysical mechanism. Numerous proteins, including other deubiquitinases, ubiquitin ligases, DNA repair and endocytosis machinery, and epigenetic regulators are functionally dependent on ubiquitin binding23. Developing selective inhibitors that attenuate ubiquitin binding is an effective strategy for USP7 inhibition. Our studies demonstrate the feasibility of this approach, which may have broader applications for inhibiting other classes of ubiquitin-binding proteins. [1] \n\nDrug resistance is a well-known phenomenon leading to a reduction in the effectiveness of pharmaceutical treatments. Resistance to chemotherapeutic agents can involve various intrinsic cellular processes including drug efflux, increased resistance to apoptosis, increased DNA damage repair capabilities in response to platinum salts or other DNA-damaging drugs, drug inactivation, drug target alteration, epithelial-mesenchymal transition (EMT), inherent cell heterogeneity, epigenetic effects, or any combination of these mechanisms. Deubiquitinating enzymes (DUBs) reverse ubiquitination of target proteins, maintaining a balance between ubiquitination and deubiquitination of proteins to maintain cell homeostasis. Increasing evidence supports an association of altered DUB activity with development of several cancers. Thus, DUBs are promising candidates for targeted drug development. In this review, we outline the involvement of DUBs, particularly ubiquitin-specific proteases, and their roles in drug resistance in different types of cancer. We also review potential small molecule DUB inhibitors that can be used as drugs for cancer treatment.[2] GNE‑6640 is a selective, non‑covalent USP7 inhibitor that binds ≈12 Å from the catalytic cysteine, attenuating ubiquitin binding. It preferentially inhibits USP7 activity toward ubiquitin moieties with free Lys48 side chains, leading to protracted depolymerization of K48‑linked polyubiquitin chains. The compound was developed via NMR‑based fragment screening and structure‑based design. It exhibits synergistic effects with PIM kinase inhibitors and DNA‑damaging chemotherapeutic agents. A somatic loss‑of‑function mutation D305G in USP7 has been found in acute lymphoblastic leukemia patients, underscoring the clinical relevance of targeting this site.[1] |
Solubility Data
| Solubility (In Vitro) |
|
|||
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 3.0268 mL | 15.1341 mL | 30.2682 mL | |
| 5 mM | 0.6054 mL | 3.0268 mL | 6.0536 mL | |
| 10 mM | 0.3027 mL | 1.5134 mL | 3.0268 mL |