Physicochemical Properties
| Molecular Formula | C24H34N6S |
| Molecular Weight | 438.6 |
| Exact Mass | 438.256 |
| CAS # | 1241537-79-0 |
| PubChem CID | 46898309 |
| Appearance | White to off-white solid powder |
| LogP | 4.7 |
| Hydrogen Bond Donor Count | 0 |
| Hydrogen Bond Acceptor Count | 6 |
| Rotatable Bond Count | 5 |
| Heavy Atom Count | 31 |
| Complexity | 566 |
| Defined Atom Stereocenter Count | 0 |
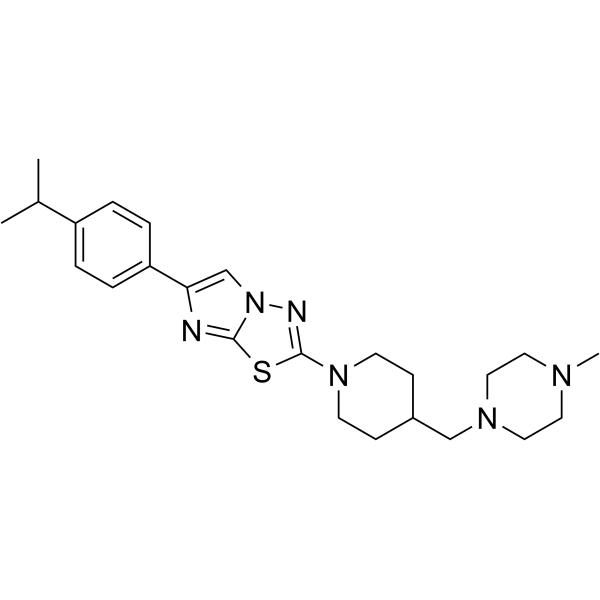
| SMILES | CC(C)C1=CC=C(C2=CN3C(SC(N4CCC(CN5CCN(C)CC5)CC4)=N3)=N2)C=C1 |
| InChi Key | RFHSWNJYHRAOMP-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C24H34N6S/c1-18(2)20-4-6-21(7-5-20)22-17-30-23(25-22)31-24(26-30)29-10-8-19(9-11-29)16-28-14-12-27(3)13-15-28/h4-7,17-19H,8-16H2,1-3H3 |
| Chemical Name | 2-[4-[(4-methylpiperazin-1-yl)methyl]piperidin-1-yl]-6-(4-propan-2-ylphenyl)imidazo[2,1-b][1,3,4]thiadiazole |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
Fer kinase and its cancer cell-specific variant FerT (kinase domain, KD) Dissociation constant (Kd) of E260 binding to Fer/FerT KD: 0.85 µM (measured by microscale thermophoresis)[1] |
| ln Vitro |
E260 is a Fer and FerT inhibitor that selectively generates metabolic stress in cancer cells by causing mitochondrial malfunction and deformation and activating energy-consuming autophagy, ultimately lowering cellular ATP levels. To demonstrate that E260 directly targets Fer and FerT, in vitro kinase tests were performed utilizing purified kinase domain (KD)-containing fragments of these enzymes. This investigation indicates a direct inhibitory action of E260 on this domain, demonstrating a substantial drop in the autophosphorylation level of Fer/FerT KD upon incubation with ATP and increasing concentrations of E260. Furthermore, computational examination of E260 docking in the simulated complete Fer protein demonstrated that the highest scoring binding mode of E260 to Fer lay within the ATP-binding pocket of enzyme KD. To estimate the dissociation constant (Kd) of E260 from Fer/FerT KD, microscale thermophoresis (MST) assays were performed using increasing concentrations of E260. This research confirmed the direct binding of E260 to the Fer/FerT KD and established the Kd to be 0.85 µM. To assess the effect of E260 micellar formulations on Fer in cancerous cells, immunoprecipitation of kinases was done in untreated and E260-treated SW620 CC cells. When given to metastatic grade IV SW620 CC cells (serum deprived for 16 hours and treated with 3 mM H2O2 to activate Fer), E260 displayed an inhibitory effect on Fer kinase activity, which was reflected in the suppression of the autophosphorylation activity of the enzyme. To describe the effect of E260 on malignant cells, metastatic SW620 cells were treated with E260 and then analyzed for vitality. The commencement of death was detected in E260-treated cells with EC50 values of 400 nM after 24 hours and 300 nM after 48 hours. E260 exhibits an EC50 of 3.2 µM after 72 hours of treatment of non-metastatic PANC-1 cells originating from primary pancreatic ductal cancer. Furthermore, the maximal amount of death in these cells was around 70% after 72 hours of treatment with 4 µM E260. In contrast, metastatic ductal carcinoma cell line SU.86.86 was demonstrated to be more susceptible to E260, with an EC50 of 1.1 µM after 72 hours of treatment, and 2 µM E260 resulted in 100% fatal levels [1]. E260 inhibits auto-phosphorylation of Fer/FerT KD in vitro, with decreasing phosphorylation levels observed with increasing concentrations of E260[1] E260 induces mitochondrial deformation, loss of mitochondrial membrane potential (MMP), and decreased complex I activity in colon cancer (CC) cells (SW620), but not in normal human fibroblasts (Hfb) or epithelial cells (CCD841CoN)[1] E260 reduces cellular ATP levels by 35–40% in SW620 cells after 12 h treatment, preceding cell death[1] E260 induces autophagy in SW620 cells, evidenced by accumulation of autophagosomes and increased LC3II/LC3I ratio[1] E260 dissociates Fer from PARP-1 in SW620 nuclei, leading to PARP-1 activation and increased poly-ADP-ribosylation[1] E260 induces necrotic death in SW620 cells, as shown by HMGB1 release, EthD-III uptake, and Annexin V/PI staining, without apoptotic markers[1] E260 exhibits selective cytotoxicity: EC50 = 400 nM (24 h) and 300 nM (48 h) in SW620 cells; no significant death in Hfb or CCD841CoN cells up to 5 µM[1] E260 downregulates hexokinase II (HK II) expression and mitochondrial association in SW620 cells under high glucose conditions[1] |
| ln Vivo |
In vivo xenograft progression is inhibited by E260. In mice, the pharmacokinetic (PK) profile of E260 was established. E260 is efficiently dispersed in animal tissues, as evidenced by its T1/2 in blood of 175 minutes and distribution volume of 4244 mL/kg. Swine Pancreatic SW620 cells were injected subcutaneously into immunocompromised "nude" mice in order to assess the effect of E260 on tumor growth. During the course of the trial, the administration of E260 significantly slowed the progression of the tumor, and after 22 days of treatment, the mean tumor volume had decreased by ten times. E260's in vivo anticancer effect was further demonstrated by treating mice with xenografts made of SW48 cells, and tracking the development of tumors was done. When compared to control treatment groups, the progression of the tumor was halted five or six times in mice administered E260 [1]. E260 (25 or 50 mg/kg, IP, twice daily for 22 days) significantly attenuates SW620 and SW48 xenograft tumor progression in immunocompromised mice fed a restricted diet[1] Tumor volume reduction was 10-fold for SW620 and 5–6-fold for SW48 compared to controls[1] Histopathology revealed necrotic and non-vascularized tumor tissue in E260-treated mice, unlike viable vascularized tumors in controls[1] No significant body weight loss or toxicity in treated animals[1] Under ad libitum feeding (high glucose), tumor suppression was reduced to 2.5-fold[1] |
| Enzyme Assay |
Recombinant Fer/FerT kinase domain (KD) protein was expressed in bacterial cells and purified using affinity resin and dialysis[1] The KD protein was incubated with ATP and increasing concentrations of E260 in kinase buffer for 1 h at room temperature[1] Samples were analyzed by SDS-PAGE and Western blot using anti-phosphotyrosine and anti-Fer antibodies to assess auto-phosphorylation inhibition[1] Microscale thermophoresis (MST) was used to measure binding affinity: labeled Fer KD was incubated with E260 in binding buffer, and Kd was calculated by nonlinear regression[1] |
| Cell Assay |
Colon cancer cell lines (SW620, SW48, HCT116) and normal cells (Hfb, CCD841CoN) were cultured in minimal essential medium (MEM)[1] Cells were seeded in 96-well plates and treated with E260 at various concentrations for specified times[1] Viability was assessed using MultiTox-Fluor or XTT assays according to manufacturer protocols[1] For mitochondrial membrane potential (MMP) measurement, cells were stained with TMRE and analyzed by microscopy or ELISA[1] Autophagy was assessed by TEM visualization of autophagosomes and immunostaining for LC3[1] HMGB1 release was quantified in culture medium using an ELISA kit[1] Necrosis was confirmed by EthD-III uptake and Annexin V/PI staining[1] Western blot analyses were performed for phospho-proteins, LC3, PARP-1, and glycolytic enzymes[1] |
| Animal Protocol |
Immunocompromised nude mice were inoculated subcutaneously with SW620 or SW48 colon cancer cells[1] Mice were fed a restricted diet (3 g/day per mouse) starting 2 days before inoculation to lower blood glucose[1] E260 was formulated in Cremophor-based nanomicelles and administered intraperitoneally at 25 or 50 mg/kg twice daily for 22 days[1] Control mice received empty micelles[1] Tumor volume was measured regularly, and mice were sacrificed at day 22 for histopathological analysis[1] Blood samples were collected for toxicology and pharmacokinetic analysis[1] |
| ADME/Pharmacokinetics |
E260 exhibited a half-life (T1/2) of 175 min in mouse blood following intraperitoneal administration[1] Volume of distribution was 4244 mL/kg, indicating efficient tissue distribution[1] |
| Toxicity/Toxicokinetics |
No significant toxicity was observed in mice treated with E260 at 25 or 50 mg/kg twice daily for 22 days[1] No body weight loss, abnormal blood chemistry (liver/kidney markers, electrolytes), or histopathological abnormalities in major organs (heart, liver, kidneys)[1] |
| References |
[1]. A novel Fer/FerT targeting compound selectively evokes metabolic stress and necrotic death in malignant cells. Nat Commun. 2017 Oct 16;8(1):940. |
| Additional Infomation |
E260 is a novel low-molecular-weight synthetic inhibitor of Fer and FerT kinases[1] It selectively induces metabolic stress, mitochondrial dysfunction, autophagy, and PARP-1 activation in malignant cells, leading to necrotic death[1] E260 is more effective against high-grade/metastatic cancer cells compared to low-grade cells[1] Its efficacy is influenced by extracellular glucose levels, with high glucose delaying but not preventing cell death[1] |
Solubility Data
| Solubility (In Vitro) | DMSO : ~1 mg/mL (~2.28 mM) |
| Solubility (In Vivo) |
Solubility in Formulation 1: ≥ 0.1 mg/mL (0.23 mM) (saturation unknown) in 10% DMSO + 40% PEG300 + 5% Tween80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 1.0 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: ≥ 0.1 mg/mL (0.23 mM) (saturation unknown) in 10% DMSO + 90% (20% SBE-β-CD in Saline) (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 1.0 mg/mL clear DMSO stock solution to 900 μL of 20% SBE-β-CD physiological saline solution and mix evenly. Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Solubility in Formulation 3: ≥ 0.1 mg/mL (0.23 mM) (saturation unknown) in 10% DMSO + 90% Corn Oil (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 1.0 mg/mL clear DMSO stock solution to 900 μL of corn oil and mix evenly. Solubility in Formulation 4: 22 mg/mL (50.16 mM) in 0.5% CMC-Na/saline water (add these co-solvents sequentially from left to right, and one by one), suspension solution; with ultrasonication. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 2.2800 mL | 11.3999 mL | 22.7998 mL | |
| 5 mM | 0.4560 mL | 2.2800 mL | 4.5600 mL | |
| 10 mM | 0.2280 mL | 1.1400 mL | 2.2800 mL |