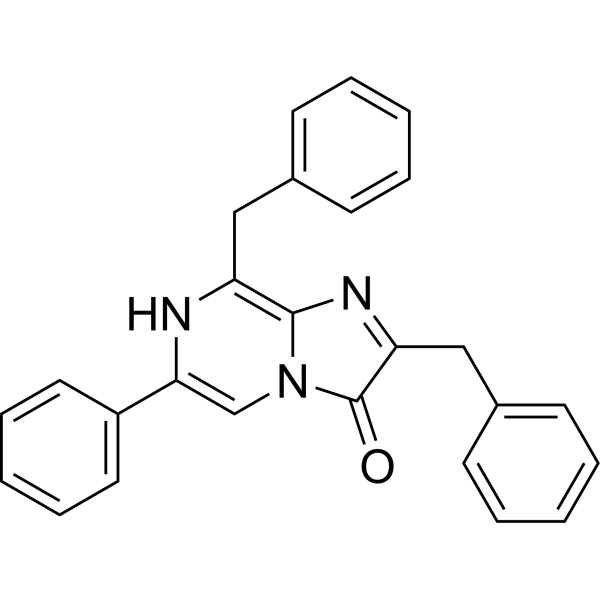
Physicochemical Properties
| Molecular Formula | C26H21N3O |
| Molecular Weight | 391.464445829391 |
| Exact Mass | 391.168 |
| CAS # | 70217-82-2 |
| Related CAS # | Coelenteramine 400a hydrochloride;2320429-05-6 |
| PubChem CID | 135750016 |
| Appearance | Brown to dark brown solid powder |
| Density | 1.2±0.1 g/cm3 |
| Boiling Point | 551.4±60.0 °C at 760 mmHg |
| Flash Point | 287.3±32.9 °C |
| Vapour Pressure | 0.0±1.5 mmHg at 25°C |
| Index of Refraction | 1.662 |
| LogP | 4.84 |
| Hydrogen Bond Donor Count | 1 |
| Hydrogen Bond Acceptor Count | 3 |
| Rotatable Bond Count | 5 |
| Heavy Atom Count | 30 |
| Complexity | 524 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | OC1=C(CC2C=CC=CC=2)N=C2C(CC3C=CC=CC=3)=NC(C3C=CC=CC=3)=CN21 |
| InChi Key | XNNYOKUWNAAJQW-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C26H21N3O/c30-26-23(17-20-12-6-2-7-13-20)28-25-22(16-19-10-4-1-5-11-19)27-24(18-29(25)26)21-14-8-3-9-15-21/h1-15,18,30H,16-17H2 |
| Chemical Name | 2,8-dibenzyl-6-phenylimidazo[1,2-a]pyrazin-3-ol |
| Synonyms | Coelenterazine 400a; 70217-82-2; Coelenterazine 400 a; Coelenteramine 400a; BlueSyn C; 2,8-dibenzyl-6-phenylimidazo[1,2-a]pyrazin-3(7H)-one; 2,8-dibenzyl-6-phenyl-7H-imidazo[1,2-a]pyrazin-3-one; 2,8-dibenzyl-6-phenylimidazo[1,2-a]pyrazin-3-ol; |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets | Luminescent enzyme substrate |
| ln Vitro | Replacement of the methylene group at the C-8 position with an extended electronic conjugation is a new promising method to develop red-shifted coelenterazine derivatives. In this paper, we have described an oxygen-containing coelenterazine derivative with a significant red-shifted (63 nm) bioluminescence signal maximum relative to coelenterazine 400a (DeepBlueC™, 1). In cell imaging, the sulfur-containing coelenterazine derivative displayed a significantly (1.77 ± 0.09; P ≤ 0.01) higher luminescence signal compared to coelenterazine 400a and the oxygen-containing coelenterazine derivative exhibited a slightly (0.74 ± 0.08; P ≤ 0.05) lower luminescence signal. It is beneficial to understand further the underlying mechanisms of bioluminescence.[3] |
| Cell Assay |
Relative quantum yield (RQY) and kinetics of in vitro luminescence[3] The RQY study and kinetic analysis were performed using an IVIS Kinetic which consisted of a cooled charge-coupled device (CCD) camera mounted on a light-tight specimen chamber (dark box), a camera controller, a camera cooling system, and controlled using a computer. The data are represented as pseudocolor images (in photons/s/cm2/scr) of light intensity (blue—least intense, red—most intense) superimposed over the grayscale reference images. Circular specified regions of interest (ROIs) were drawnon the areas, and the light output were quantified as the total number of photons emitted per second using Living Image software. To determine the appropriate unsaturated amount of substrate, 10 μL of coelenterazine (CTZ) derivatives 1–3 between 1 and 100 μmol/L and 90 μL of either DMSO or Rluc enzyme at a final concentration of 15 nmol/L (bioluminescence) were used. The RQY and reaction kinetics was determined by mixing 10 μL of coelenterazine (CTZ) derivatives (final concentration of 5 μmol/L) with 90 μL of either DMSO or Rluc enzyme (15 nmol/L) onto wells of 96-well black plates to prevent light reflection from well to well. Luminescent signals were measured immediately after mixing and monitored over a period of 25–30 min (luminescence had almost decayed to near-background levels) using the IVIS. Light output was recorded every 5 min with an exposure time of 30 s for chemiluminescence and every 1 min with an exposuretime of 5 s in the first 15 min for bioluminescence. The collected data was analyzed by employing the Prism 5.0 GraphPad software to compute the total light output. As a corresponding blank control, Tris–HCl buffer was added instead of coelenterazine (CTZ) derivatives solution under the same conditions. All assays were performed in triplicate. Assay for luminescence activity in live cell[3] ES-2 cells (human ovarian cancers cell line) expressing Rluc were supplied by BioDiagnosis. The ES-2 cells were cultured in Dulbecco's modified Eagle's medium (DMEM; high glucose with l-glutamine) containing 10% fetal bovine serum (FBS) and 0.5 μg/mL puromycin at 37 °C in a humidified atmosphere in a 5% CO2 incubator.coelenterazine (CTZ) derivatives was dissolved in ethanol to make a 1 mmol/L stock solution, and diluted with Tris–HCl buffer to gradient concentration (5, 10, 20, 40, 60, 80, 100 μmol/L). The ES-2-Rluc cells were grown in black 96-well plates (4 × 104 cells per well). After a 24-h incubation period, the medium was removed. Then cells were washed with Tris–HCl buffer twice and treated with 100 μL of a series of concentrations of coelenterazine (CTZ) derivatives solutions (ranging from 0 to 100 μmol/L). Bioluminescent signals were then immediately determined using the IVIS. Light output was recorded every 1 min with an exposure time of 30 s until luminescence almost decayed to near-background levels. Luminescent signal (photons per second) for each well was measured and plotted as average values. All experiments were performed in triplicate. |
| References |
[1]. Bisdeoxycoelenterazine derivatives for improvement of bioluminescence resonance energy transfer assays. J Am Chem Soc. 2007 Oct 3;129(39):11900-1. [2]. The BRET2/arrestin assay in stable recombinant cells: a platform to screen for compounds that interact with G protein-coupled receptors (GPCRS). J Recept Signal Transduct Res. Feb-Nov 2002;22(1-4):533-41. [3]. Luminescence of coelenterazine derivatives with C-8 extended electronic conjugation. Chinese Chemical Letters, 2016, 27(4): 550-554. |
| Additional Infomation |
In conclusion, we have described an oxygen-containing CTZ derivative 2 and it exhibited a more significant red-shifted (63 nm) bioluminescence signal maximum relative to coelenterazine 400a, while it had lower quantum yield. The results of this study appear to support our hypothesis that the introduction of oxygen heteroatom at the C-8 position could also produce a bathochromic effect. While a possible mechanism of the CTZ luminescence reaction has been proposed (Fig. S2 in Supporting information), the enzymatic recognition mechanism of the Rluc system is not yet fully understood. We may surmise that it is due to an efficient extension of p-electron conjugation, electronegativity, or H-bonds of oxygen at the C-8 position. We also developed a new measurement method through an IVIS Kinetic equipped with a CCD to examine RQY. The RQY of CTZ derivative 3 for chemiluminescence is in good agreement with previously reported value. To get a genuine and correct RQY in bioluminescence, we employed commercially available purified Rluc enzyme in place of the rough cytosolic extract to avoid enzyme-independent luminescence. Compounds 2 and 3 showed lower quantum yield that is quite different from previously reported value. However, we discovered compound 3 had a higher affinity to Rluc than coelenterazine 400a. In cell imaging, compound 3 also displayed a higher luminescence signal. In a word, compounds 2 and 3 are promising bright red-shifted CTZ derivatives that provide a novel approach to improve the luminescence properties of CTZ analogs. More significantly, it is beneficial to understand the underlying mechanisms of Rluc bioluminescence upon reacting with coelenterazine.[3] In BRET2 (Bioluminescence Resonance Energy Transfer), a Renilla luciferase (RLuc) is used as the donor protein, while a Green Fluorescent Protein (GFP2) is used as the acceptor protein. In the presence of the cell permeable substrate DeepBlueC, RLuc emits blue light at 395 nm. If the GFP2 is brought into close proximity to RLuc via a specific biomolecular interaction, the GFP2 will absorb the blue light energy and reemit green light at 510nm. BRET2 signals are therefore easily determined by measuring the ratio of green over blue light (510/395nm) using appropriate dual channel luminometry instruments (e.g., Fusion Universal Microplate Analyzer, Packard BioScience). Since no light source is required for BRET2 assays, the technology does not suffer from high fluorescent background or photobleaching, the common problems associated with standard FRET-based assays. Using BRET2, we developed a generic G Protein-Coupled Receptor (GPCR) assay based on the observation that activation of the majority of GPCRs by agonists leads to the interaction of beta-arrestin (a protein that is involved in receptor desensitization and sequestration) with the receptor. We established a cell line stably expressing the GFP2:beta-arrestin 2 fusion protein, and showed that it can be used to monitor the activation of various transiently expressed GPCRs, in BRET2/arrestin assays. In addition, using the HEK 293/GFP2:beta-arrestin 2 cell line as a recipient, we generated a double-stable line co-expressing the vasopressin 2 receptor (V2R) fused to RLuc (V2R:RLuc) and used it for the pharmacological characterization of compounds in BRET2/arrestin assays. This approach yields genuine pharmacology and supports the BRET2/arrestin assay as a tool that can be used with recombinant cell lines to characterize ligand-GPCR interactions which can be applied to ligand identification for orphan receptors.[2] |
Solubility Data
| Solubility (In Vitro) | May dissolve in DMSO (in most cases), if not, try other solvents such as H2O, Ethanol, or DMF with a minute amount of products to avoid loss of samples |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 2.5545 mL | 12.7727 mL | 25.5454 mL | |
| 5 mM | 0.5109 mL | 2.5545 mL | 5.1091 mL | |
| 10 mM | 0.2555 mL | 1.2773 mL | 2.5545 mL |