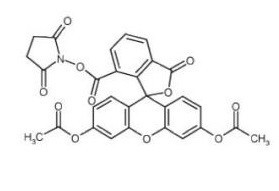
CFSE [5(6)-Carboxyfluorescein diacetate succinimidyl ester; CFDA-SE; 5(6)-CFDA N-succinmidyl ester] is a novel, a cell-permeable and amine-reactive fluorescent dye which has been widely used to track cell division by covalently coupling, via its succinimidyl group, to intracellular molecules such as lysine residues and other amine sources. Specifically, it is used to monitor distinct generations of proliferating cells by dye dilution. It is non-fluorescent in the parent form until the acetate groups are cleaved by intracellular esterases to produce the highly fluorescent fluorophore.
Physicochemical Properties
| Molecular Formula | C₂₉H₁₉NO₁₁ |
| Molecular Weight | 557.46 |
| Exact Mass | 557.095 |
| Elemental Analysis | C, 62.48; H, 3.44; N, 2.51; O, 31.57 |
| CAS # | 150347-59-4 |
| Appearance | Off-white to yellow solid powder |
| Density | 1.6±0.1 g/cm3 |
| Boiling Point | 757.9±70.0 °C at 760 mmHg |
| Melting Point | 152-154ºC(lit.) |
| Flash Point | 412.2±35.7 °C |
| Vapour Pressure | 0.0±2.6 mmHg at 25°C |
| Index of Refraction | 1.701 |
| LogP | 0.5 |
| InChi Key | JGPOSNWWINVNFV-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C29H19NO11/c1-14(31)37-17-4-7-20-23(12-17)39-24-13-18(38-15(2)32)5-8-21(24)29(20)22-11-16(3-6-19(22)28(36)40-29)27(35)41-30-25(33)9-10-26(30)34/h3-8,11-13H,9-10H2,1-2H3 |
| Chemical Name | (2,5-dioxopyrrolidin-1-yl) 3',6'-diacetyloxy-1-oxospiro[2-benzofuran-3,9'-xanthene]-5-carboxylate |
| Synonyms | 5(6-Carboxyfluorescein diacetate succinimidyl ester; CFDA-SE; 5(6-CFDA N-succinmidyl ester; |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month Note: Please store this product in a sealed and protected environment (e.g. under nitrogen), avoid exposure to moisture and light. |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| Targets |
Fluorescent dye Preparation of CFDA-SE working solution 1.1 Prepare stock solution: it is recommended to dissolve 1 milligram CFDA-SE in 0.1794 mL DMSO to obtain 10 mM CFDA-SE. Note: Store the stock solution at -20℃ or -80℃ in the dark and avoid repeated freezing. 1.2 Preparation of CFDA-SE working solution. Dilute stock solution in serum-free cell culture. Note: Please adjust the concentration of CFDA-SE working solution based on your specific needs. Cell staining 2.1 Suspended cells: Centrifuge at 1000 g for 3-5 minutes at 4°C and discard the supernatant. Use PBS to wash twice, for five minutes each time. Adherent cells: Discard the cell culture media, apply trypsin to separate the cells and produce a single cell suspension. Centrifuge at 1000 g for 3-5 minutes at 4°C and discard the supernatant. Use PBS to wash twice, for five minutes each time. 2.2 Add 1 mL CFDA-SE working solution and mix for 30 minutes. 2.3 Centrifuge at 400 g for 3-4 minutes at 4°C. 2.4 Wash the cells twice with PBS, five minutes/each time. 2.5 Resuspend the cells in serum-free medium or in PBS, and detect using fluorescence microscopy or flow cytometer. Cell proliferation labeling: CFSE can be used to label cells in vitro for proliferation assays. It freely diffuses into cells as a non - fluorescent form. Inside the cell, it is cleaved by cellular esterases, and the succinimidyl group irreversibly binds to intracellular amino groups, forming a fluorescent conjugate that remains in the cell. When cells divide, the fluorescent conjugate is evenly distributed to two daughter cells, and the fluorescence intensity of daughter cells is half of that of the parent cell. Flow cytometry can be used to detect the series - halved fluorescence intensity of CFSE - labeled cell proliferation populations [1] |
| ln Vitro |
Preparation of CFDA-SE working solution 1.1 Prepare stock solution: it is recommended to dissolve 1 milligram CFDA-SE in 0.1794 mL DMSO to obtain 10 mM CFDA-SE. Note: Store the stock solution at -20℃ or -80℃ in the dark and avoid repeated freezing. 1.2 Preparation of CFDA-SE working solution. Dilute stock solution in serum-free cell culture. Note: Please adjust the concentration of CFDA-SE working solution based on your specific needs. Cell staining 2.1 Suspended cells: Centrifuge at 1000 g for 3-5 minutes at 4°C and discard the supernatant. Use PBS to wash twice, for five minutes each time. Adherent cells: Discard the cell culture media, apply trypsin to separate the cells and produce a single cell suspension. Centrifuge at 1000 g for 3-5 minutes at 4°C and discard the supernatant. Use PBS to wash twice, for five minutes each time. 2.2 Add 1 mL CFDA-SE working solution and mix for 30 minutes. 2.3 Centrifuge at 400 g for 3-4 minutes at 4°C. 2.4 Wash the cells twice with PBS, five minutes/each time. 2.5 Resuspend the cells in serum-free medium or in PBS, and detect using fluorescence microscopy or flow cytometer. Cell proliferation labeling: CFSE can be used to label cells in vitro for proliferation assays. It freely diffuses into cells as a non - fluorescent form. Inside the cell, it is cleaved by cellular esterases, and the succinimidyl group irreversibly binds to intracellular amino groups, forming a fluorescent conjugate that remains in the cell. When cells divide, the fluorescent conjugate is evenly distributed to two daughter cells, and the fluorescence intensity of daughter cells is half of that of the parent cell. Flow cytometry can be used to detect the series - halved fluorescence intensity of CFSE - labeled cell proliferation populations [1] CFSE (carboxyfluorescein succinimidyl ester) is used as an intracellular fluorescent dye for tracking cell proliferation. In cell proliferation assays, CFSE fluorescence intensity (FI) within cells decreases over time due to natural decay and protein turnover, even in the absence of cell division. This decay exhibits a biphasic pattern and can be mathematically modeled. Experimental data from unstimulated PBMC cultures from two healthy donors showed a time-dependent decrease in mean CFSE FI over 160 hours. [1] The study compared multiple mathematical models to describe the label decay kinetics. Models incorporating multiple loss rate mechanisms (e.g., separate rates for different fluorescent conjugates) or a single time-dependent decay rate (e.g., Gompertz decay model, \( \frac{dx}{dt} = -c(x - x_a)e^{-kt} \) ) provided better fits to the observed CFSE decay data compared to a simple exponential decay model. [1] CFSE is used to track proliferating cells by flow cytometry, allowing discrimination of up to eight or more successive cell divisions. It has been applied to study division-related phenotypic and functional changes during differentiation of B cells, T cells, and hematopoietic precursor cells. It can also be combined with immunophenotyping using antibodies conjugated to PE, PerCP, or PE/Cy5. [2] CFSE staining intensity is linear with respect to concentration, and fluorescence decays by about 60% in the first 1–2 days in non-dividing cells, then stabilizes for weeks to months. [2] The dye has been used to examine slowly dividing glioblastoma cancer cells, erythroid progenitor proliferation, antigen-specific T cell clones, and neuroantigen-specific T cells in multiple sclerosis. [2] |
| ln Vivo |
CFDA-SE is an intracellular and green fluorescent dye that may be used to detect strain colonization in vivo. Method: For determine strain colonization in the intestine. 1. Strains are resuspended in PBS to obtain a cell density of 108 CFU/mL. 2. CFDA-SE (1 mM; 10 μL; 20 min; 37°C; dark/protect from light) is added to 1 mL of bacterial suspension. 3. Centrifuge (13000 g, 10 min, 4°C) and wash the mixture for 3 times using sterile PBS to remove excess CFDA-SE. 4. Strains are resuspended in sterile PBS (108 CFU/mL) and 1 mL of bacterial suspension is administered to SD rats. Day 1 and 3 after gavage, the fluorescence imaging is performed on anesthetized rats who are then sacrificed and different intestinal sections are taken for imaging. 5. The fluorescence imaging is performed using an IVIS Lumina III Smart Imaging System. - Cell tracking: In vivo, CFSE can remain in viable cells for several weeks at measurable concentrations, regardless of cell type or activation state. It can be used to track the division and proliferation of cells in the body, providing uniform labeling with little adverse effects on the cell's intracellular machinery [2] CFSE-labeled cells can be adoptively transferred in vivo (e.g., intravenous injection in mice) and tracked for several months. It has been used to study B cell division in the absence of T cell division, alloresponses in mixed lymphocyte reactions, and graft-versus-host response modulation. [2] |
| Cell Assay |
The technique described in this unit uses the intracellular fluorescent label carboxyfluorescein diacetate succinimidyl ester (CFSE) to track proliferating cells. Covalently bound CFSE is divided equally between daughter cells, allowing discrimination of successive rounds of cell division. The technique is applicable to in vitro cell division, as well as to in vivo division of adoptively transferred cells and can resolve eight or more successive generations. CFSE is long lived, permitting analysis for several months after cell transfer, and has the same spectral characteristics as fluorescein, so monoclonal antibodies conjugated to phycoerythrin or other compatible fluorochromes may be used to immunophenotype the dividing cells. In addition, information is given on a second-generation dye, Cell Trace Violet (CTV), excited by 405-nm blue laser light. CTV is chemically related to CFSE, but allows the 488-nm line of the Argon laser to be used for other probes[2].
Cell labeling and proliferation detection: Prepare cell suspensions, add an appropriate amount of CFSE solution, and incubate at an appropriate temperature for a certain period to allow CFSE to enter the cells. Then, wash the cells to remove unbound CFSE. Culture the labeled cells under appropriate conditions. At different time points, use flow cytometry to measure the fluorescence intensity of cells. According to the change of fluorescence intensity, analyze cell proliferation, division and other conditions [1] The standard procedure for CFSE labeling of cells involves using its precursor, carboxyfluorescein diacetate succinimidyl ester (CFDA-SE). CFDA-SE, due to its lipophilicity, passively diffuses across cell membranes and is taken up by cells. Once inside the cell, intracellular esterases (specifically, acetylesterase is hypothesized) catalyze the hydrolysis of its acetate esters, converting it into the highly fluorescent and membrane-impermeable CFSE. The succinimidyl ester moiety of CFSE then reacts with intracellular amine groups on proteins, forming stable fluorescent conjugates (e.g., CF-R2) that are retained, and unstable conjugates (e.g., CF-R1) that are lost or degraded. After initial exposure and staining, the cell culture is flushed to remove excess label. [1] For the decay study, peripheral blood mononuclear cells (PBMCs) from two donors were stained with CFSE following the standard procedure but were not stimulated to divide. This ensured any observed fluorescence loss was due to natural decay processes. Cells were measured at 24 distinct time points over 160 hours in triplicate using flow cytometry, and the mean total fluorescence intensity (FI) of each sample was recorded. [1] Cells are resuspended in PBS/0.1% BSA at 5 × 10⁷ cells/ml. CFSE is added to a final concentration of 10 µM and incubated for 10 minutes at 37°C. Staining is quenched with ice-cold RPMI 1640/10% FBS for 5 minutes on ice. Cells are washed three times in culture or injection medium. For proliferation assays, cells are cultured under appropriate conditions or adoptively transferred. Harvested cells can be stained with antibodies for immunophenotyping and analyzed by flow cytometry with excitation at 488 nm and emission collected with a 525-nm band-pass filter. [2] For improved uniformity, CFSE can be diluted to 20 µM in PBS/0.1% BSA and added to an equal volume of 2× concentrated cell suspension. [2] |
| Animal Protocol |
For adoptive transfer, CFSE-labeled cells (e.g., 1 × 10⁷ to 5 × 10⁷ cells per mouse) are injected intravenously via the tail vein in mice or rats. Cells are tracked in lymphoid organs such as the spleen, and analyzed by flow cytometry at various time points post-transfer. [2] |
| ADME/Pharmacokinetics |
CFSE fluorescence intensity decreases by approximately 60% within the first 1–2 days in non-dividing cells due to catabolism of labile bound components, after which it remains stable for several weeks to months. [2] The dye is long-lived in vivo, allowing tracking for several months post-transfer. [2] |
| Toxicity/Toxicokinetics |
High concentrations of CFSE can impact cell viability and division. Optimal staining should give a 3–3.5 log increase in fluorescence intensity relative to unstained cells. Including protein (0.1% BSA) during staining improves viability and reduces aggregation. Staining at lower temperatures (ice or room temperature) may also improve viability. [2] CFSE can modify expression of some cell surface markers or lower staining by some monoclonal antibodies, possibly due to steric hindrance. [2] |
| References |
[1]. Quantifying CFSE Label Decay in Flow Cytometry Data. Appl Math Lett. 2013 May 1;26(5):571-577. [2]. Flow cytometric analysis of cell division by dilution of CFSE and related dyes. Curr Protoc Cytom. 2013;Chapter 9:Unit9.11. [3]. Prostaglandin E3 attenuates macrophage-associated inflammation and prostate tumour growth by modulating polarization. J Cell Mol Med. 2021 Jun;25(12):5586-5601. [4]. Long-lived pancreatic ductal adenocarcinoma slice cultures enable precise study of the immune microenvironment. Oncoimmunology. 2017; 6(7): e1333210. |
| Additional Infomation |
We developed a series of models for the label decay in cell proliferation assays when the intracellular dye carboxyfluorescein succinimidyl ester (CFSE) is used as a staining agent. Data collected from two healthy patients were used to validate the models and to compare the models with the Akiake Information Criteria. The distinguishing features of multiple decay rates in the data are readily characterized and explained via time dependent decay models such as the logistic and Gompertz models.[1] Pancreatic ductal adenocarcinoma (PDA) remains a deadly disease that is rarely cured, despite many recent successes with immunotherapy for other malignancies. As the human disease is heavily infiltrated by effector T cells, we postulated that accurately modeling the PDA immune microenvironment would allow us to study mechanisms of immunosuppression that could be overcome for therapeutic benefit. Using viable precision-cut slices from fresh PDA, we developed an organotypic culture system for this purpose. We confirmed that cultured slices maintain their baseline morphology, surface area, and microenvironment after at least 6 d in culture, and demonstrated slice survival by MTT assay and by immunohistochemistry staining with Ki-67 and cleaved-Caspase-3 antibodies. Immune cells, including T cells (CD3+, CD8+, and FOXP3+) and macrophages (CD68+, CD163+ and HLA-DR+), as well as stromal myofibroblasts (αSMA+) were present throughout the culture period. Global profiling of the PDA proteome before and after 6 d slice culture indicated that the majority of the immunological proteins identified remain stable during the culture process. Cytotoxic effects of drug treatment (staurosporine, STS and cycloheximide, CHX) on PDA slices culture confirmed that this system can be used to assess functional response and cell survival following drug treatment in both a treatment time- and dose-dependent manner. Using multicolor immunofluorescence, we stained live slices for both cancer cells (EpCAM+) and immune cells (CD11b+ and CD8+). Finally, we confirmed that autologous CFSE-labeled splenocytes readily migrate into co-cultured tumor slices. Thus, our present study demonstrates the potential to use tumor slice cultures to study the immune microenvironment of PDA.[4] CFSE is a colorless, non - fluorescent, non - polar molecule. It has a succinimidyl group that can specifically bind to cells and a carboxyfluorescein diacetate group that can be hydrolyzed non - enzymatically. It is a good live - cell marker. The use of CFSE to label cells can replace traditional methods such as morphological observation and 3H - TdR incorporation method in immunological experiments, which can improve the research level and expand the application of new immunological techniques [2] CFSE is the active fluorescent form generated intracellularly from its precursor, CFDA-SE. It is a vital tool for mapping cellular division histories in proliferation assays. Upon covalent binding to long-lived intracellular proteins (forming CF-R2), the fluorescent label is retained in viable cells for several weeks, allowing long-term tracking. The loss of fluorescence over time (label decay) is attributed to natural decay of the dye and turnover of the proteins to which it is conjugated. [1] The primary application discussed is the use of CFSE in flow cytometry-based cell proliferation assays. The serial dilution of CFSE fluorescence with each cell division creates a correlation between measured fluorescence intensity and the number of divisions a cell has undergone. Accurately modeling the natural decay of CFSE fluorescence is crucial for correctly estimating cell proliferation and death rates from such data. [1] Mathematical models explored in the literature to quantify CFSE label decay include systems of ordinary differential equations representing the conversion and decay of different intracellular fluorescent species (CFDA-SE, CFSE, CF-R1, CF-R2), as well as simplified single-equation models using Exponential, Logistic, or Gompertz decay kinetics. The Gompertz decay model, which features a time-dependent decay rate, was found to provide a significantly better fit to the observed biphasic decay data than a simple exponential model. [1] CFSE (carboxyfluorescein diacetate succinimidyl ester) is a cell-permeant fluorescein-based dye that covalently binds to intracellular amines, allowing equal distribution between daughter cells upon division. It is excited at 488 nm and emits at ~525 nm, compatible with fluorescein filter sets. [2] A second-generation dye, Cell Trace Violet (CTV), excited at 405 nm, offers similar division tracking capability and frees the 488-nm channel for other probes. [2] CFSE data can be used with modeling software (e.g., ModFit) to calculate precursor frequency and proliferation indices. [2] |
Solubility Data
| Solubility (In Vitro) | DMSO : ~50 mg/mL (~89.69 mM) |
| Solubility (In Vivo) |
Solubility in Formulation 1: ≥ 2.08 mg/mL (3.73 mM) (saturation unknown) in 10% DMSO + 40% PEG300 + 5% Tween80 + 45% Saline (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 20.8 mg/mL clear DMSO stock solution to 400 μL PEG300 and mix evenly; then add 50 μL Tween-80 to the above solution and mix evenly; then add 450 μL normal saline to adjust the volume to 1 mL. Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH₂ O to obtain a clear solution. Solubility in Formulation 2: ≥ 2.08 mg/mL (3.73 mM) (saturation unknown) in 10% DMSO + 90% Corn Oil (add these co-solvents sequentially from left to right, and one by one), clear solution. For example, if 1 mL of working solution is to be prepared, you can add 100 μL of 20.8 mg/mL clear DMSO stock solution to 900 μL of corn oil and mix evenly. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 1.7939 mL | 8.9693 mL | 17.9385 mL | |
| 5 mM | 0.3588 mL | 1.7939 mL | 3.5877 mL | |
| 10 mM | 0.1794 mL | 0.8969 mL | 1.7939 mL |