1M7 is a reagent for RNA SHAPE-MaP experiments that enables analysis of RNA secondary structures at single nucleotide resolution in<70 seconds. SHAPE chemistry performed with 1M7 accurately reports the known structure of the RNase P specificity domain under native conditions. 1M7 reactivity detects nucleotides constrained both by base pairing and by idiosyncratic, noncanonical tertiary interactions. SHAPE chemistry enables very precise analysis of the differences between two structures, such as Mg2+-dependent tertiary interactions. 1M7 is easily handled in the laboratory and enables analysis of large RNA structures at single nucleotide resolution in less than 70 s.
Physicochemical Properties
| Molecular Formula | C9H6N2O5 |
| Molecular Weight | 222.15434 |
| Exact Mass | 222.027 |
| Elemental Analysis | C, 48.66; H, 2.72; N, 12.61; O, 36.01 |
| CAS # | 73043-80-8 |
| Related CAS # | 73043-80-8; |
| PubChem CID | 12535373 |
| Appearance | Light yellow to yellow solid powder |
| Density | 1.6±0.1 g/cm3 |
| Boiling Point | 395.4±44.0 °C at 760 mmHg |
| Flash Point | 192.9±28.4 °C |
| Vapour Pressure | 0.0±0.9 mmHg at 25°C |
| Index of Refraction | 1.623 |
| LogP | 0.4 |
| Hydrogen Bond Donor Count | 0 |
| Hydrogen Bond Acceptor Count | 5 |
| Rotatable Bond Count | 0 |
| Heavy Atom Count | 16 |
| Complexity | 350 |
| Defined Atom Stereocenter Count | 0 |
| SMILES | O=C(O1)N(C)C2=CC([N+]([O-])=O)=CC=C2C1=O |
| InChi Key | MULNCJWAVSDEKJ-UHFFFAOYSA-N |
| InChi Code | InChI=1S/C9H6N2O5/c1-10-7-4-5(11(14)15)2-3-6(7)8(12)16-9(10)13/h2-4H,1H3 |
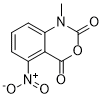
| Chemical Name | 1-Methyl-7-nitro-2H-3,1-benzoxazine-2,4(1H)-dione |
| Synonyms | 1M7; 1-methyl-7-nitro-3,1-benzoxazine-2,4-dione |
| HS Tariff Code | 2934.99.9001 |
| Storage |
Powder-20°C 3 years 4°C 2 years In solvent -80°C 6 months -20°C 1 month Note: This product requires protection from light (avoid light exposure) during transportation and storage. |
| Shipping Condition | Room temperature (This product is stable at ambient temperature for a few days during ordinary shipping and time spent in Customs) |
Biological Activity
| ln Vitro | Selective 2‘-hydroxyl acylation analyzed by primer extension (SHAPE) chemistry allows local nucleotide flexibility to be quantitatively assessed at single nucleotide resolution in any RNA. SHAPE chemistry exploits structure-based gating of the nucleophilic reactivity of the ribose 2‘-hydroxyl group by the extent to which a nucleotide is constrained or flexible. SHAPE chemistry was developed using N-methylisatoic anhydride (NMIA), which is only moderately electrophilic and requires tens of minutes to form ribose 2‘-O-adducts. Here, we design and evaluate a significantly more useful, fast-acting, reagent for SHAPE chemistry. Introduction of a nitro group para to the reactive carbonyl to form 1-methyl-7-nitroisatoic anhydride (1M7) yields a reagent that both reacts significantly more rapidly with RNA to form 2‘-O-adducts and is also more labile toward advantageous, self-limiting, hydrolysis. With 1M7, the single nucleotide resolution interrogation of the RNA structure is complete in 70 s. SHAPE analysis performed with 1M7 accurately reports the secondary and tertiary structure of the RNase P specificity domain and allows the secondary structure of this RNA to be predicted with up to 91% accuracy [1]. |
| Enzyme Assay | Many biological processes are RNA-mediated, but higher-order structures for most RNAs are unknown, which makes it difficult to understand how RNA structure governs function. Here we describe selective 2'-hydroxyl acylation analyzed by primer extension and mutational profiling (SHAPE-MaP) that makes possible de novo and large-scale identification of RNA functional motifs. Sites of 2'-hydroxyl acylation by SHAPE are encoded as noncomplementary nucleotides during cDNA synthesis, as measured by massively parallel sequencing. SHAPE-MaP-guided modeling identified greater than 90% of accepted base pairs in complex RNAs of known structure, and we used it to define a new model for the HIV-1 RNA genome. The HIV-1 model contains all known structured motifs and previously unknown elements, including experimentally validated pseudoknots. SHAPE-MaP yields accurate and high-resolution secondary-structure models, enables analysis of low-abundance RNAs, disentangles sequence polymorphisms in single experiments and will ultimately democratize RNA-structure analysis [2]. |
| Cell Assay | SHAPE-MaP is unique among RNA structure probing strategies in that it both measures flexibility at single-nucleotide resolution and quantifies the uncertainties in these measurements. We report a straightforward analytical framework that incorporates these uncertainties to allow detection of RNA structural differences between any two states, and we use it here to detect RNA-protein interactions in healthy mouse trophoblast stem cells. We validate this approach by analysis of three model cytoplasmic and nuclear ribonucleoprotein complexes, in 2 min in-cell probing experiments. In contrast, data produced by alternative in-cell SHAPE probing methods correlate poorly (r = 0.2) with those generated by SHAPE-MaP and do not yield accurate signals for RNA-protein interactions. We then examine RNA-protein and RNA-substrate interactions in the RNase MRP complex and, by comparing in-cell interaction sites with disease-associated mutations, characterize these noncoding mutations in terms of molecular phenotype. Together, these results reveal that SHAPE-MaP can define true interaction sites and infer RNA functions under native cellular conditions with limited preexisting knowledge of the proteins or RNAs involved.[3] |
| References |
[1]. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J Am Chem Soc . 2007 Apr 11;129(14):4144-5. doi: 10.1021/ja0704028. [2]. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat Methods . 2014 Sep;11(9):959-65. doi: 10.1038/nmeth.3029. [3]. Detection of RNA-Protein interactions in living cells with SHAPE. Biochemistry . 2015 Nov 24;54(46):6867-75. doi: 10.1021/acs.biochem.5b00977. |
| Additional Infomation | 1-methyl-7-nitroisatoic anhydride is a 3,1-benzoxazin-1,4-dione having an N-methyl substituent and a nitro group at the 7-position. It is a benzoxazine and a C-nitro compound. |
Solubility Data
| Solubility (In Vitro) | DMSO : ~50 mg/mL (~225.07 mM) |
| Solubility (In Vivo) |
Note: Listed below are some common formulations that may be used to formulate products with low water solubility (e.g. < 1 mg/mL), you may test these formulations using a minute amount of products to avoid loss of samples. Injection Formulations (e.g. IP/IV/IM/SC) Injection Formulation 1: DMSO : Tween 80: Saline = 10 : 5 : 85 (i.e. 100 μL DMSO stock solution → 50 μL Tween 80 → 850 μL Saline) *Preparation of saline: Dissolve 0.9 g of sodium chloride in 100 mL ddH ₂ O to obtain a clear solution. Injection Formulation 2: DMSO : PEG300 :Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL DMSO → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Injection Formulation 3: DMSO : Corn oil = 10 : 90 (i.e. 100 μL DMSO → 900 μL Corn oil) Example: Take the Injection Formulation 3 (DMSO : Corn oil = 10 : 90) as an example, if 1 mL of 2.5 mg/mL working solution is to be prepared, you can take 100 μL 25 mg/mL DMSO stock solution and add to 900 μL corn oil, mix well to obtain a clear or suspension solution (2.5 mg/mL, ready for use in animals). Injection Formulation 4: DMSO : 20% SBE-β-CD in saline = 10 : 90 [i.e. 100 μL DMSO → 900 μL (20% SBE-β-CD in saline)] *Preparation of 20% SBE-β-CD in Saline (4°C,1 week): Dissolve 2 g SBE-β-CD in 10 mL saline to obtain a clear solution. Injection Formulation 5: 2-Hydroxypropyl-β-cyclodextrin : Saline = 50 : 50 (i.e. 500 μL 2-Hydroxypropyl-β-cyclodextrin → 500 μL Saline) Injection Formulation 6: DMSO : PEG300 : castor oil : Saline = 5 : 10 : 20 : 65 (i.e. 50 μL DMSO → 100 μLPEG300 → 200 μL castor oil → 650 μL Saline) Injection Formulation 7: Ethanol : Cremophor : Saline = 10: 10 : 80 (i.e. 100 μL Ethanol → 100 μL Cremophor → 800 μL Saline) Injection Formulation 8: Dissolve in Cremophor/Ethanol (50 : 50), then diluted by Saline Injection Formulation 9: EtOH : Corn oil = 10 : 90 (i.e. 100 μL EtOH → 900 μL Corn oil) Injection Formulation 10: EtOH : PEG300:Tween 80 : Saline = 10 : 40 : 5 : 45 (i.e. 100 μL EtOH → 400 μLPEG300 → 50 μL Tween 80 → 450 μL Saline) Oral Formulations Oral Formulation 1: Suspend in 0.5% CMC Na (carboxymethylcellulose sodium) Oral Formulation 2: Suspend in 0.5% Carboxymethyl cellulose Example: Take the Oral Formulation 1 (Suspend in 0.5% CMC Na) as an example, if 100 mL of 2.5 mg/mL working solution is to be prepared, you can first prepare 0.5% CMC Na solution by measuring 0.5 g CMC Na and dissolve it in 100 mL ddH2O to obtain a clear solution; then add 250 mg of the product to 100 mL 0.5% CMC Na solution, to make the suspension solution (2.5 mg/mL, ready for use in animals). Oral Formulation 3: Dissolved in PEG400 Oral Formulation 4: Suspend in 0.2% Carboxymethyl cellulose Oral Formulation 5: Dissolve in 0.25% Tween 80 and 0.5% Carboxymethyl cellulose Oral Formulation 6: Mixing with food powders Note: Please be aware that the above formulations are for reference only. InvivoChem strongly recommends customers to read literature methods/protocols carefully before determining which formulation you should use for in vivo studies, as different compounds have different solubility properties and have to be formulated differently. (Please use freshly prepared in vivo formulations for optimal results.) |
| Preparing Stock Solutions | 1 mg | 5 mg | 10 mg | |
| 1 mM | 4.5015 mL | 22.5073 mL | 45.0146 mL | |
| 5 mM | 0.9003 mL | 4.5015 mL | 9.0029 mL | |
| 10 mM | 0.4501 mL | 2.2507 mL | 4.5015 mL |